
A new influenza virus had previously been isolated from little yellow-shouldered bats (Sturnira lilium) in Guatemala. Three of 316 rectal swabs were positive when tested by a pan-influenza polymerase chain reaction assay. Viral sequences were also detected in liver, intestine, lung, and kidney tissues, suggestive of viral replication and not passage of ingested material through the intestinal tract. Analysis of the viral genome sequence revealed that A/little yellow-shouldered bat/Guatemala/164/2009 (H17N10) is significantly diverged from all known influenza viruses.
When the same PCR approach was used to screen 114 rectal swabs from 18 different species of bats captured in Peru, a single flat-faced fruit bat (Artibeus planirostris) was positive. Viral sequences were also detected in liver, intestine, and spleen tissues from the same bat. Comparison of the sequences of all 8 genome RNA segments with those of the H17N10 Guatemalan isolate revealed sufficient divergence to justify naming it a new HA and NA subtype, A/flat-faced bat/Peru/033/2010 (H18N11).
Comparison of the nucleotide sequences of bat influenza A viruses from Peru and Guatemala with other influenza viruses leads to two amazing conclusions. First, 7 of the 8 viral RNAs of the bat influenza A viruses group separately from the RNAs of all other known influenza viruses. Second, the RNA sequences encoding four proteins, PB2, PB1, PA and NA, display greater genetic diversity than in all non-bat influenza virus sequences combined. The implication is that New World bats harbor a diverse pool of influenza viruses.
The H17 and H18 HA RNA sequences are, in contrast, far more related to known influenza virus HA and NA sequences. The implication of this observation is clear: some time after the bat and non-bat influenza A viruses diverged, a reassortment event occurred that introduced the HA of a non-bat influenza A virus into the genome of a bat influenza A virus.
Serological studies have revealed widespread circulation of these two new influenza viruses in bats. Sera from 55 of 110 (50%) Peruvian bats representing 13 different species were positive for antibodies against the viral HA or NA proteins. Twenty-one of these samples were positive for antibodies against both viral glycoproteins, while 30 were positive only for anti-HA18 antibodies and 4 were positive for only anti-N11 antibodies. These observations suggest that some bats are infected with reassortant viruses carrying the H18 or N11 genes. A study of sera from 8 different species of Guatemalan bats revealed antibodies to the H17 HA protein in 86 of 228 sera (38%).
A number of human viruses, such as SARS-coronavirus and Nipah and Hendra viruses, are known to have originated in bats. Can bat influenza A viruses infect humans and serve as a source of future pandemic strains? The answer to this question is not known, but the two new bat viruses cannot infect human cell lines in culture. However, it is possible that acquisition of other (e.g. avian or swine) influenza virus genes by reassortment could produce a virus with bat influenza virus genes that is capable of infection humans. The pathogenic and pandemic potential of such viruses is unknown. A first step to answering this question would be to determine if human populations with contact with bats have antibodies to the two new bat influenza A viruses.
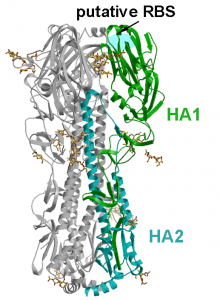
The cell receptor for all known influenza A viruses is the carbohydrate molecule known as sialic acid The cell receptor for the two new bat influenza A viruses is not known, but it is clearly not sialic acid, a conclusion reached by studying the crystal structures and binding properties of the H17 (paper one and two) and H18 HA (illustrated) molecules. Furthermore, the crystal structures of the N10 (paper one and two) and N11 proteins reveal that their substrate cannot be sialic acid (the function of the influenza A virus NA is to remove sialic acids from the cell surface, allowing newly synthesized virions to move away from the cell). For this reason the N10 and N11 proteins are called ‘NA-like’.
Bats also harbor many other kinds of viruses, including hepatitis B viruses, Marburg virus, hepaciviruses, pegiviruses, paramyxoviruses, coronaviruses, and many more. They also contain parasites – specifically, malaria parasites. For more information, listen to these podcast episodes:
Hedging our bats (TWiV 258)
More bats out of hell (TWiP 62)
Hepaciviruses and pegiviruses in bats and rodents (TWiV 231)
Bats out of hell (TWiV 183)
Going to bat for flu research (TWiV 173)
Matt’s bats (TWiV 65)

What other possible implications are there of the incredible diversity in bat PB1, PB2, PA, and NA genes? Could influenza A viruses in bats predate the circulation of these viruses in wild birds? Could the different structure of sialic acid-binding HAs have any effect on capsid stability if you put one on a bat influenza virus? That might be an important factor in zoonotic potential, which would presumably require a reassortment leading to acquisition of a sialic acid-binding HA.
Fascinating!
You may follow our special issue on “Viruses and Bats” at
http://www.mdpi.com/journal/viruses/special_issues/viruses_bats
And
so
> Could influenza A viruses in bats predate the circulation of these
viruses in wild birds?
influenza inner segments don’t mutate at the amino-acid level in
mallards.
In mammals, including bats, it mutates “away”, so mallard flu seems to
be
the original.
other than the Gustemala virus, the Peru virus has high AT-content,
as we usually only see after decades,centuries of evolution in humans,
swine,horses. So I speculate that there could be a reservoir in
Piccaries,Tapirs,
http://www.flutrackers.com/forum/showthread.php?t=212800
Pingback: Bat SARS-like coronavirus, influenza and NEIDL: ‘What’s New on Virology Blog?’ - The Global Dispatch