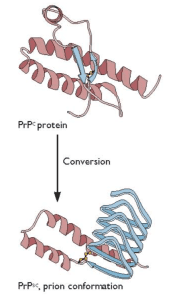

Spongiform encephalopathies are neurodegenerative diseases caused by misfolding of normal cellular prion proteins. Human spongiform encephalopathies are placed into three groups: infectious, familial or genetic, and sporadic, distinguished by how the disease is acquired initially. In all cases, the pathogenic protein is the host-encoded PrPC protein with an altered conformation, called PrPsc. In the simplest case, PrPSc converts normal PrPC protein into more copies of the pathogenic form (illustrated).
The structure of the normal PrPC protein, solved some time ago, revealed that it is largely alpha-helical with little beta-strand content. The structure of PrPSc protein has been elusive, because it forms aggregates and amyloid fibrils. It has been suggested that the PrPSc protein has more beta-strand content than the normal protein, but how this property would lead to prion replication was unknown. Clearly solving the structure of prion protein was needed to fully understand the biology of this unusual pathogen.
The structure of PrPSc protein has now been solved by cryo-electron microscopy and image reconstruction (link to paper). The protein was purified from transgenic mice programmed to produce a form of PrPSc protein that is not anchored to the cell membrane, and which is also underglycosylated. The protein causes disease in mice but is more homogeneous and forms fibrillar plaques, allowing gentler purification methods.
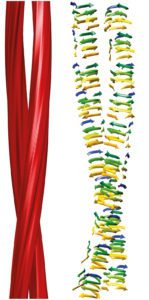

The authors note that the molecular interactions that control prion templating, including hydrogen-bonding, charge and hydrophobic interactions, aromatic stacking, and steric constraints, also play roles in DNA replication.
The structure of PrPSc protein provides a mechanism for prion replication by incorporation of additional molecules into a growing beta-solenoid. I wonder if incorporation into fibrils is the sole driving force for converting PrPCprotein into PrPSc, or if PrPC is conformationally altered before it ever encounters a growing fibril.

To know more about prions, which book or reading is recommended?