
The first crystallographic X-ray structure of a virus was that of tomato bushy stunt virus in 1977, followed by structures of the animal viruses poliovirus and rhinovirus in 1985. Since then hundreds of viral structures have been determined. Scientists know that the coordinates for these structures can be found at the Protein Data Bank. However, it is not a straightforward task to produce a complete viral capsid from those data, and non-scientists searching for a pre-made complete virus image may find the site daunting.
VIPERdb solves these problems with its collection of icosahedral virus capsid structures. For the scientist, there are comprehensive data from structural and computational analyses of different viruses. For those who simply want to see what a virus looks like, there are high quality renderings for visual exploration. In these images, all viral capsids are placed in a single icosahedral orientation convention, making it easy to compare different structures. The web site includes powerful search utilities, links to other relevant databases, background information on virus capsid structure, and useful database interface tools.
I have always found VIPERdb an extremely useful tool for making virus images for research or teaching. I asked my colleagues at VIPERdb to answer questions about their resource for the readers of virology blog.
Why did you establish ViperDB?
VIPER, in its early stage, was started because the icosahedral virus coordinates deposited in the Protein Data Bank (PDB) were not organized in such a way that a non expert could easily generate a single virus particle, or a particular oligomer of subunits (pentamer, trimer, dimer). It was clear that if it took an expert in structural virology 3-4 hours to determine the appropriate symmetry matrices to create a single complete particle, members of the virology community interested in using the structures were frustrated and probably not using the information. Therefore all the icosahedral virus coordinates in the protein data bank were systematically reoriented into the newly defined VIPER convention and made publicly available (we note that the pdb now stores virus coordinates in this convention, although they do not provide all the analysis features VIPER has. The PDB has been very cooperative and continues to be interactive).
Preliminary analysis of virus capsids in terms of the contacts at the various interfaces was started when Vijay Reddy joined Jack Johnson’s group in 1992-1993 at Purdue University. The adhoc approaches to virus capsid analysis culminated into robust and uniform analysis of virus structures, when the group moved to The Scripps Research Institute (TSRI) and started a collaboration with Charles Brooks III, and joined the NIH Research Resource Center for the Development of Multiscale Modeling Tools for Structural Biology (MMTSB) in 1997-1998. VIPER evolved eventually into VIPERdb, when the storage flat files where moved into a MySQL database, and continues to evolve as new and sophisticated features are added. Careful manual curation on a per-entry basis is another big feature of the VIPERdb site. A lot of time is spent making sure each new capsid entered is in accordance with the VIPER convention and standards.
A lot of virus capsid symmetry and quasi-symmetry knowledge came from years of Johnson’s group expertise. Brooks group provided the know-how to uniform analysis of virus capsids using CHARMM simulation and analysis computer program. Vijay Reddy acted as a bridge between the two groups, directing the analysis of capsids and design of web tools with the help and assistance of a number of post doctoral fellows and summer students (Mauricio Carrillo-Tripp, Craig M. Shepherd, Sangita Venkataraman, Gabriel Lander, Padmaja Natarajan, Komath Damodaran, Ryan Morton, Kevin Li, Brian Okerberg, Ian A. Borelli), making significant contributions over the years, resulting in ten publications so far. The VIPERdb site continues to be supported by the MMTSB resource.
Who is the target audience for ViperDB?
Our target audience is virologists who are interested in using virus particle structure information for their studies. With the explosion of infectious clones and the ability to produce virus like particles in bacuolovirus systems and E. coli, there has been an interest in making rational mutants based on the structure. Now we also have bioinformatics people interested in this resource because of the large amount of data on protein-protein interfaces. We also see the site as an educational resource, with convenient generation of quasi-equivalent surface lattices and generation of galleries of viruses.
Would a non-scientist be able to navigate ViperDB, if they wanted to simply download a capsid image?
We have invested a lot of effort trying to make the site easy to navigate and to have a user friendly interface. Each virus entry has its own ‘Info Page’ containing the corresponding illustrations, visualization tools, and analysis data (annotations), and there are several ways to access them, such as PDB ID and drop-down boxes in the top menu and several options in the ‘Find-a-virus’ page. A quick way for a non scientist to get to a capsid image is to browse the database using the ‘Family Index’ tool located under the ‘Data’ top menu. This application interface shows links to all entries in the database grouped by family. Another way to get an ‘images only’ page is the ‘Galley Maker’ tool, under the ‘Utilities’ top menu. Here, the user can create a gallery of virus renderings by setting several options. Leaving all the default values will create a page with an scaled topology image of every single entry in VIPERdb.
Do you have plans to include non-icosahedral structures, e.g. viral glycoproteins?
We have certainly considered this for the future. Right now we are working on completing the latest stage of the site, where we want to take it to become a web application, in which case the user can use and interact with the data in several dynamic ways and do their own analysis.
Can anyone use images made on ViperDB? How should they acknowledge you?
Absolutely. Anyone can use any of the images and data available on the site. We only ask for the corresponding acknowledgement, either by creating a link back to us (in case of a web related work) or by including a reference to the published paper in the journal Nucleic Acid Research 2009, D436:37 (in case of a printed work).
Do you have any new features planned for ViperDB?
Yes, some of them are already sketched in there and some others are already functional. As we mentioned before, we are trying to make the site a web application. This can be seen already in the ‘Info Page’, specifically in the 3D IAU (Icosahedral Asymmetric Unit) and Phi-Psi Explorer Tools tabs. We make use of Jmol to dynamically explore several aspects of the three dimensional structure of the proteins in relation to the different capsid symmetry axes, in the case of the former, and the Google Maps API (Application Programming Interface) in a novel and unique way to explore protein-protein interactions and extent of sequence conservation within families in a visual way, in case of the latter. A more detailed explanation on the Phi-Psi Maps can be found on the Phi-Psi Explorer tab of any Info Page.
The structure alignment tools are another couple of working features of the renovated site. We implemented the Top-Match and the STRAP applications providing exceptional structure analysis tools to the site. Both have their own set of specific characteristics and we take good advantage of them.
We also are developing our own API to interact with the database through a more straight forward programmatic way, accessed with regular HTTP calls. This should help create different, and more specialized, branches off the VIPERdb site in the future.
There is a new ‘Discussion Forums’ section where we would like to build a community of users. We believe that as the number of individuals that participate at all levels grow, everybody will benefit from it. On this line of thought, we are producing a set of “screen casts” directed at those who are not familiar with the site and its different sections and tools.
On a more ambitious note, we have started a project to build homology models of a high amount of structurally uncharacterized viruses. We are using the sequence of identified members of an specific family to build three dimensional models based on a member of the same family whose structure is known. We plan to populate the database with these models, taking the current number of entries, 264, to the hundreds or thousands. We anticipate that this will help answer open questions related to virus assembly and capsid stability, among others.
Do you have any favorite virus images?
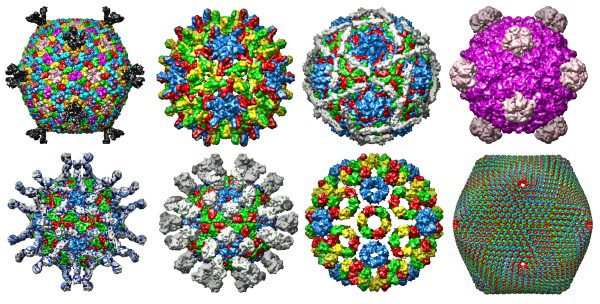
X-ray structure of the entire lipid-containing bacteriophage pm2 (2w0c), Corticoviridae
Poliovirus/Receptor Complex (1nn8) Enterovirus, Picornaviridae
Human Hepatitis B Viral Capsid (1qgt) Orthohepadnavirus, Hepadnaviridae
Foot and Mouth Disease Virus/Fab Complex (1qgc) Aphthovirus, Picornaviridae
Echovirus 12 (1upn) Enterovirus, Picornaviridae
Semliki Forest Virus (1dyl) Alphavirus, Togaviridae
Bacteriophage Φ-X 174 (2bpa) Microvirus, Microviridae
PBCV-1 Virus Capsid (1m4x) Chlorovirus, Phycodnaviridae
A gallery of these viral capsids (not to scale) is at the top of this post.
Many thanks to Mauricio Carrillo Tripp and the team at VIPERdb for explaining their site. It’s a great resource, so do head on over and give it a try. Send us your favorite virus images from VIPERdb, and we’ll post them here at virology blog!
F. K. Winkler, C. E. Schutt, S. C. Harrison, G. Bricogne (1977). Tomato bushy stunt virus at 5.5-Ã… resolution Nature, 265 (5594), 509-513 DOI: 10.1038/265509a0

Very nice (and useful post) – glad to hear that FMDV numbers among their favorites!
It(He) is very friendly! Me happy much of obtaining this page. Someone podria to say to me if it(he,she) exists algun link to recreate the empty capsida of a virus?
It(He) is very friendly! Me happy much of obtaining this page. Someone podria to say to me if it(he,she) exists algun link to recreate the empty capsida of a virus?
VIPERdb crashed my browser twice when I tried to view structures
Thank you for sharing the effort and cooperation that made VIPERdb possible. We hope the db is expandable on comodity hardware, and open source. There is much that could be done with good resources to expand it. We see it as a “pilot plant” for other projects, as Evergreen is an open source bibliographic tool.
Pingback: ¡Recapitulemos! | CientÃficos del siglo XX y su legado