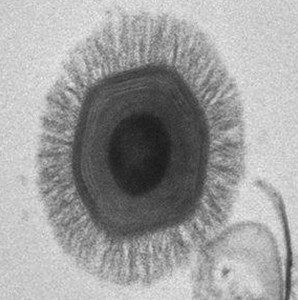

To search for giant viruses in humans, respiratory samples from 196 Tunisian patients with community-acquired pneumonia were co-cultured with the amoebae Acanthamoeba polyphaga. One sample, a bronchial aspirate, yielded a new Megavirus that was given the catchy name LBA111. The patient from which LBA111 was isolated had been hospitalized with fever, cough, shortness of breath, and bloody sputum. Serum from this patient, but not from those of 50 healthy blood donors, reacted with LBA111 proteins, indicating suggesting that this individual had been infected with the virus.
Mimivirus LBA111 is not likely to be a contaminant introduced from the laboratory in which it was isolated, because its genome sequence is original. The LBA111 genome is double-stranded DNA 1,230,522 base pairs in length, mostly closely related to the genome of Megavirus chilensis, a giant virus isolated off the coast of Chile. In addition to other differences, the LBA111 genome encodes two tRNAs (histidine and cysteine) not present in the genome of M. chilensis.
While these findings indicate an association of Mimivirus LBA111 with human pneumonia, they do not prove that this virus is the causative agent of human disease. However, the possibility that Mimivirus causes human disease makes sense. Mimiviruses are present in soil and water where they multiply in amoebae. A known agent of human respiratory disease, Legionella pneumophila, also colonizes amoebae. Antibodies to Mimivirus, as well as Mimivirus DNA, have been found in patients with pneumonia. Mimivirus should therefore be added to the list of agents that should be considered in patients with pneumonia.

Pingback: The largest viral genome from a human | Viral Bioinformatics Resource Center
You say, ” Serum from this patient, but not from those of 50 healthy blood donors, reacted with LBA111 proteins, indicating that this individual had likely been infected with the virus.” I’m not sure that follows. The humoral response to LBA111 proteins suggests that this individual had been exposed to those proteins, but if LBA111 infected an ameba that was present in this individual, wouldn’t that be more parsimonious (since we have no evidence that Mimiviruses infect humans)? It is interesting to consider whether detection of Mimiviruses and Megaviruses could be a marker of amebic infection.
What if it is preying on the human microbiota? Then they are “friends” rather then “enemies” (health scientists have a weird non-biological normative thinking about super-organisms, isn’t it?) Quid bono?
That’s a reasonable point. I’ll change the wording to ‘suggest’ instead of ‘indicating’.
Possibly. You are right, LBA111 might not be an enemy, but that’s our first reaction when such associations are found.
It might be useful to look for antibodies to amobae to support this idea.
Pingback: Links 7/18/13 | Mike the Mad Biologist
In case you haven’t seen it yet, there’s a new record-breaking virus: http://www.nature.com/news/giant-viruses-open-pandora-s-box-1.13410