Back in January 2020, I wrote about a deletion that appeared in the genome of SARS-CoV during the middle and late phase of the epidemic (link to article). A similar deletion has now been detected in the genome of SARS-CoV-2 (link to article).
The deletion in the SARS-CoV genome occurred in a region encoding a protein called Orf8. It was initially suggested that the 29 nucleotide deletion was somehow involved in adaptation of SARS-CoV to humans. This hypothesis was disproven in subsequent experiments which demonstrated that the 29 nucleotide deletion decreases viral replication in a number of different cell types. In other words, this virus has reduced fitness compared with a virus containing a full Orf8.
SARS-CoV-2 viruses with a 382 nucleotide deletion, which encompasses almost the entire open reading frame of Orf8, have now been isolated from eight hospitalized patients in Singapore (image below). These viruses appear to have been circulating in Singapore for at least four weeks.
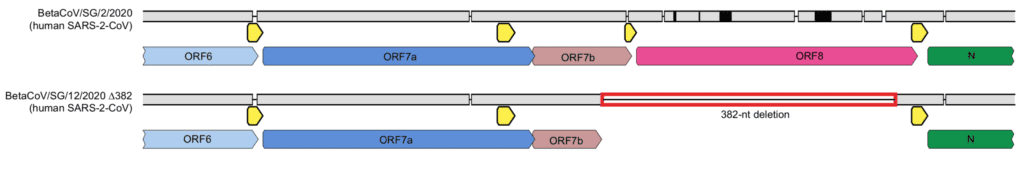

The authors of the study suggest that the deletion might lead to reduced virulence of SARS-CoV-2. This hypothesis is based on the reduced fitness of SARS-CoV lacking Orf8 in cell culture. In the article that I published here on 23 January 2020, I wrote “The genome sequence of 2019-CoV (the name at the time for SARS-CoV-2) shows that Orf8 is intact. If it is not lost during subsequent virus circulation in humans, the outbreak could be more severe.”
So far the pandemic caused by SARS-CoV-2 has been quite severe with a crude fatality rate globally of ~6%. Whether or not the Orf8 deletion will reduce virulence of the virus in humans is unknown. By analogy with SARS-CoV, the Orf8 deletion likely reduces the fitness of SARS-CoV-2 and therefore the mutant virus might not be able to compete with viruses with the intact gene.
Orf8 deleted SARS-CoV-2 viruses have not yet been reported outside of Singapore. It will be important to look carefully at this region of the genome in future virus isolates to determine if this altered virus is capable of global spread, and if so, whether it is accompanied by reduced virulence.

Pingback: A 382 nucleotide deletion in the genome of SARS-CoV-2 – Virology Hub
https://www.bloomberg.com/news/articles/2020-04-11/coronavirus-vaccine-could-be-ready-in-six-months-times
Now there is a claim that the COVID-19 vaccine could be out in September or October. If this is true.
A vaccine against the coronavirus could be ready by September, according to a scientist leading one of Britain’s most advanced teams.
Sarah Gilbert, professor of vaccinology at Oxford University, told The Times on Saturday that she is “80% confident†the vaccine would work, and could be ready by September. Experts have warned the public that vaccines typically take years to develop, and one for the coronavirus could take between 12 to 18 months at best.
In the case of the Oxford team, however, “it’s not just a hunch, and as every week goes by we have more data to look at,†Gilbert told the London newspaper.
Gilbert’s team is one of dozens worldwide working on a vaccine and is the most advanced in Britain, she told the Times. As the country looks set to begin its fourth week under lockdown, a vaccine could be fundamental in easing the measures and returning to normal life. Gilbert said human trials are due to start in the next two weeks.
Her remarks came as the death toll from the virus pushed past 100,000 globally. On Friday, the U.K. reported 980 fatalities, taking the total count from the virus to 8,958, and the government has repeatedly pleaded with the public to obey lockdown rules during the long Easter holiday weekend. As Prime Minister Boris Johnson begins his recovery after a spell in intensive care, Patrick Vallance, the government’s chief scientific adviser, warned he expects the number of deaths to increase for “a few weeks†yet.
Manufacturing the millions of vaccine doses necessary could take months. Gilbert said she’s in discussions with the British government about funding, and starting production before the final results are in, allowing the public to access the vaccine immediately if it proves to work. She said success by the autumn was “just about possible if everything goes perfectly.â€
great work dear keep it up Covid 19 – Your Defense Against It
At the end of this very interesting discussion about the 382 nt deletion in the ORF8 of SARS-CoV-2 you state that “Orf8 deleted SARS-CoV-2 viruses have not yet been reported outside of Singapore”. This may longer be the case since an identical deletion in a SARS-CoV-2 sequence from Taiwan has recently been reported.