
The first non-retroviral endogenous virus described was bornavirus, a virus with a negative-stranded RNA genome. Bornaviral sequences were found in the genomes of humans, non-human primates, rodents, and elephants. Phylogenetic analyses revealed that these sequences entered the primate genome over 40 million years ago. Endogenous filovirus (ebolavirus, marburgvirus) sequences were subsequently identified in the genomes of bats, rodents, shrews, tenrecs and marsupials. Based on these analyses it was estimated that filoviruses are at least tens of millions of years old. The presence of endogenous bornavirus and filovirus sequences were subsequently confirmed and extended to 19 different vertebrate species. Endogenous hepadnaviruses probably entered the genome of the zebra finch 19 million years ago.
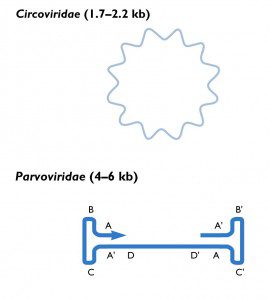
Recent additions to the endogenous virus catalog are the circoviruses and parvoviruses. The genome of circoviruses are composed of single-stranded DNA, while those of parvoviruses are linear single-stranded DNAs with base-paired ends (figure). Phylogenetic analyses of these endogenous viral sequences reveal that both virus families are 40 to 50 million years old. Examination of insect genomes has revealed endogenous viral sequences from members of the Bunyaviridae, Rhabdoviridae, Orthomyxoviridae, Reoviridae, and Flaviviridae.
With the exception of retroviruses, these endogenous viral sequences have no role in viral replication – they are accidentally integrated into host DNA. Such sequences are highly mutated and typically comprise only fragments of the viral genome, and therefore cannot give rise to infectious virus. Whether these sequences confer any biological advantage to the host is an interesting question. It is possible that some of the endogenous viral sequences are copied into RNA, or translated into protein, and could have consequences for the host. For example, it has been suggested that synthesis of the bornaviral N protein from endogenous sequences might render the host resistant to infection with bornaviruses.
How are non-retroviral genomes integrated into the host DNA? For viruses with an RNA genome, the nucleic acid must enter the nucleus (perhaps accidentally for viruses without a nuclear phase) and be converted to a DNA copy by reverse transcriptase encoded by endogenous retroviruses. Hepadnaviruses encode a reverse transcriptase which produces the genomic DNA from an RNA template. In all cases, recombination could lead to integration of viral DNA into the host chromosome.
Almost half of the human genome is made up of mobile genetic elements, which includes endogenous proviruses and other sequences derived from retroviruses such as retrotransposons, retroposons, and processed pseudogenes. It seems likely that even more diverse viral sequences lurk in cellular genomes, awaiting discovery.
Horie M, Honda T, Suzuki Y, Kobayashi Y, Daito T, Oshida T, Ikuta K, Jern P, Gojobori T, Coffin JM, & Tomonaga K (2010). Endogenous non-retroviral RNA virus elements in mammalian genomes. Nature, 463 (7277), 84-7 PMID: 20054395
Taylor DJ, Leach RW, & Bruenn J (2010). Filoviruses are ancient and integrated into mammalian genomes. BMC evolutionary biology, 10 PMID: 20569424
Belyi VA, Levine AJ, & Skalka AM (2010). Unexpected inheritance: multiple integrations of ancient bornavirus and ebolavirus/marburgvirus sequences in vertebrate genomes. PLoS pathogens, 6 (7) PMID: 20686665
Gilbert C, & Feschotte C (2010). Genomic fossils calibrate the long-term evolution of hepadnaviruses. PLoS biology, 8 (9) PMID: 20927357
Katzourakis A, & Gifford RJ (2010). Endogenous viral elements in animal genomes. PLoS genetics, 6 (11) PMID: 21124940
Belyi VA, Levine AJ, & Skalka AM (2010). Sequences from ancestral single-stranded DNA viruses in vertebrate genomes: the parvoviridae and circoviridae are more than 40 to 50 million years old. Journal of virology, 84 (23), 12458-62 PMID: 20861255

I guess there will be another debate about chicken or egg! Virologists will claim viruses exsited way before anything else on earth. But at the moment people believed viruses evolved from cells (transposon elements).
If a constant inhibitory diet has caused progressive changes in the delicate biological balance of the human being, micro-organisms have developed in the body, acting like endogenous mutagenic agents.
Virologist face many Dengerous tasks .. Wait Goat flu 2015
Don’t endogenous retroviruses replicate in the placenta when a female is pregnant? without ERV’s, a viable fetus could not be. we have evolved with the help of ERV’s which is what separates us from other primates.
There is good evidence that endogenous retroviruses of sheep play an
important role in development of the placenta. In these animals
virions are produced in the epithelia of the female reproductive
tract. Endogenous viral proteins (not virions) are produced in the
human placenta, and there is evidence from studies in cultured cells
that syncytin 1, a protein encoded by the env gene of the endogenous
retrovirus ERVWE1, is involved in the fusion of mononuclear
cytotrophoblasts to produce syncytiotrophoblasts. Whether this protein
is important for development of the placenta in humans is not known.
Pingback: The Strange Thing about Creationism - Page 28 - Religious Education Forum