
Although the role of swine as “mixing vessels” for influenza A(H1N1) viruses was established more than a decade ago, it appears that the policy makers and scientific community have underestimated it. In fact, in 1998 influenza experts proposed the establishment of surveillance in swine populations as a major part of an integrated early warning system to detect pandemic threats for humans but, to some extent, this task was overlooked.
In support of their contention, the authors compare the number of swine influenza A sequences (4,648) with those of human (46,911) and avian (41,142) influenza A viruses. They also point out that some countries, such as the United States, have devoted all of their pandemic preparedness budget ($3.8 billion for the US) towards the prevention and control of avian A(H5N1) influenza. They believe that
…in this plan, a substantial effort was dedicated to prevent and contain the foreign threat of Asian avian flu, neglecting the influenza threat that the North American swine population presents. Specifically, we believe that the aforementioned strategy ignores the swine farm and industry workers which constitute the population at higher risk of contracting and spreading the hypothetical pandemic influenza virus.
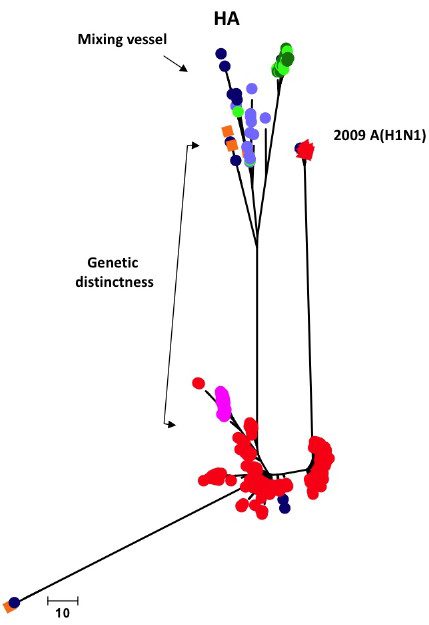
This strategy is based on the authors’ conclusions that the new influenza H1N1 viruses are genetically distinct from other H1N1 viruses that have been circulating in humans for the past 20 years. The new swine derived viruses are more similar to viruses that were transmitted from pigs to humans in Iowa, Maryland, and Wisconsin between 1991 and 2006. This conclusion is illustrated by the phylogenetic tree of HA protein sequences from H1N1 influenza viruses. Included in the analysis are H1 HA proteins from the 2009 influenza A(H1N1) virus (red triangles), earlier human (red and pink circles), swine (navy blue and purple circles), and avian (green circles) viruses. The viruses involved in pig-human interspecies transmission in Iowa, Maryland and Wisconsin, shown as orange squares, cluster with the swine viruses and the 2009 H1N1 virus.
An extensive surveillance of the genetic evolution of influenza A viruses that circulate in Mexican poultry farms has been carried out since 2002. A similar system will now be put into place on swine farms. The goal is to identify genetically distinct influenza viruses that could lead to pandemics.
G M Nava, M S Attene-Ramos, J K Ang, & M Escorcia (2009). Origins of the new influenza A(H1N1) virus: time to take action Eurosurveillance, 14 (22)

Sincere question: how would it have helped to identify the looming threat of this swine H1N1 even if we had paid more attention. As a layman, it seems to me that some of what we thought we knew about what was required for an influenza virus to adapt to humans is constantly changing.
So, how would we have know what changes to look for?
Perhaps we'd be better off working under the assumption that we can't predict which strains will manifest as a pandemic and do things that will help us against these pandemics in general. Are there such things we can do? Things that are useful regardless of whether H1N1 or H5N1 or H2N2 hits?
Great question. You are right, we cannot look at the sequence of an influenza virus and say whether or not it has the ability to infect and transmit among humans. But we might be able to see viruses evolving that have the potential to infect humans. In this sense we might have noticed that there were H1N1 viruses evolving in pigs that were very different from the current human H1n1 strains and very similar to pig viruses that had jumped into humans. Such viruses would have the potential to infect humans because the viral proteins are so different and not recognized by the immune system. It is only retrospectively do we realize how H1N1 viruses were evolving in pigs.
We can't predict which strains will extensive transmit among humans, but we can identify candidates. A key would be to identify viruses that are circulating in other animals that are antigenically diverse from those infecting humans, but which occasionally 'jump' into humans, such as the swine viruses mentioned in the post.
Pingback: H1N1 Swine Flu Update | Science News
I really do not feel that the Swine flu was underestimated. Many argue that the press and concern was over-the-top. I don't agree with this either. Although some did push the envelope a bit further than necessary. Since this viral type was similar to the catastrophe of the Spanish Flu – there was reason for concern. Also it was something new – New is always scary.
Vincent, please let me know if I'm way off on this one.
I think the “official” number of infected Americans is somewhere around 15,000 but most experts admit that you can add at least one or maybe even two zero's behind this number for the actual number of Americans infected (with mild symptoms).
Most experts are predicting huge demand for the vaccine this fall. Natural infection of the virus during the Spring offers a high level of protection to similar circulating Fall strains.
So the point I am trying to make is that maybe we should urge “would-be” vaccine recipients to get tested to see what their current “influenza antibody profile” is prior to vaccination so the people that truly need vaccine protection get the limited number of vaccinations. Of course, if the subtyping tests are considerably more expensive than the vaccine then this might be a tough stance to take.
i don't think so, but probably coz symptoms are just the same having a normal flu that's why people thought it'll go away after day and never took it as a different kind of sickness
Antibody profiling is not something that is routinely done as part of
primary health care; it will likely be difficult to convince any doc
to do it. Personally I will take the vaccine in the fall, with the
expectation that even if the virus has diverged the infection will be
milder.
Antibody profiling is not something that is routinely done as part of
primary health care; it will likely be difficult to convince any doc
to do it. Personally I will take the vaccine in the fall, with the
expectation that even if the virus has diverged the infection will be
milder.