
Two studies of newly isolated monoclonal antibodies against influenza virus suggest that the answer could be yes. The authors of one study identified human antibodies against influenza virus by phage display. In this technique, recombinant HA protein (the H5 subtype) was used to bind bacteriophage particles that bear on their surfaces variable chains of human antibodies. Ten antibodies were identified that neutralized the infectivity of H5 influenza viruses in cell culture. The antibodies also protected mice against lethal H5N1 influenza even when administered after infection.
The key result is that the monoclonal antibodies neutralize infectivity not only of H5 viruses, but also viruses of 9 other HA subtypes. There are 16 known HA subtypes, divided into two groups. The 10 subtypes neutralized by the monoclonal antibodies comprise group 1, which includes H1, H2, and H5. If another epitope can be identified that elicits neutralizing antibodies against group 2 HA subtypes, then a universal vaccine that confers life-long protection might be feasible.
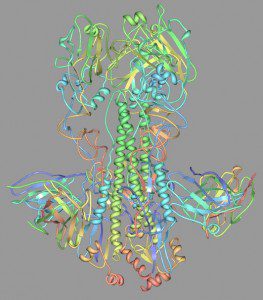
Such broadly-reacting neutralizing monoclonal antibodies have been reported previously, but they are rare. When animals are immunized with influenza virus, most of the antibodies that are produced are directed against the membrane-distal, globular head of the HA molecule (top of image). Resolution of the X-ray structure of one of the monclonal antibodies bound the the H5 HA protein revealed that the antibody binding site is a hydrophobic pocket on the stem of the HA molecule. This membrane-proximal site is probably not readily recognized by B cell receptors and therefore rarely gives rise to antibodies. The authors circumvented this problem by selecting antibodies using a soluble form of the HA protein, which is not subject to such steric constraints.
A number of significant hurdles remain to be overcome before these findings translate into an influenza vaccine. As noted above, another epitope must still be identified that elicits neutralizing antibodies against viruses of the 6 other HA types. Additional screening of human antibodies with soluble HA protein will presumably address this issue. A more difficult is how to formulate a vaccine with these epitopes. The most effective influenza vaccines are whole or split virus preparations, but how can these be prepared so that the membrane-proximal HA epitope is immunodominant? If the globular head of the HA is removed, the virus will not be infectious and cannot be propagated for vaccine production. One solution might be to produce defective particles in producer cell lines. The problem is not insurmountable, but will require some new and innovative approaches in influenza vaccine development.
These are very exciting findings, and in my opinion, bode well for a universal influenza vaccine within the next decade.
Jianhua Sui, William C Hwang, Sandra Perez, Ge Wei, Daniel Aird, Li-mei Chen, Eugenio Santelli, Boguslaw Stec, Greg Cadwell, Maryam Ali, Hongquan Wan, Akikazu Murakami, Anuradha Yammanuru, Thomas Han, Nancy J Cox, Laurie A Bankston, Ruben O Donis, Robert C Liddington, Wayne A Marasco (2009). Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses Nature Structural & Molecular Biology DOI: 10.1038/nsmb.1566
Mark Throsby, Edward van den Brink, Mandy Jongeneelen, Leo L. M. Poon, Philippe Alard, Lisette Cornelissen, Arjen Bakker, Freek Cox, Els van Deventer, Yi Guan, Jindrich Cinatl, Jan ter Meulen, Ignace Lasters, Rita Carsetti, Malik Peiris, John de Kruif, Jaap Goudsmit (2008). Heterosubtypic Neutralizing Monoclonal Antibodies Cross-Protective against H5N1 and H1N1 Recovered from Human IgM+ Memory B Cells PLoS ONE, 3 (12) DOI: 10.1371/journal.pone.0003942

Pingback: Headless HA: universal influenza vaccine?