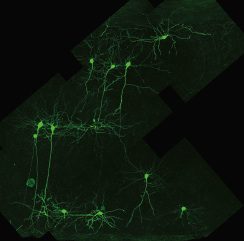

The GFP gene was first inserted into the genome of potato virus X, a plant pathogen. One to two days after inoculation of the recombinant virus onto plants, regions of the leaves fluoresced bright green when illuminated with UV light. Animal viruses followed several years later, with insertion of the GFP gene into the genome of adenovirus. Infection with the recombinant virus lead to green fluorescence in cultured mammalian cells and in living brain slices, suggesting the possibility of using this technology to trace neuronal connections. This potential was subsequently realized when recombinant viruses that produce GFP were used to trace neuronal connections in the nervous system of living animals. Proteins like GFP that emit light of different wavelengths have been discovered in different organisms, allowing the tracking of multiple proteins in virus infected cells.
More recently, fluorescent proteins have been used to visualize single virus particles in living cells. In an early validation of the technique, the gene encoding the herpes simplex virus type 1 VP26 protein, which is located on the outer surface of the capsid, was fused with the coding sequence for GFP. The fusion protein was incorporated into intranuclear capsids and mature virions, which produced green fluorescence in infected cells. By using such viruses, entry, uncoating, replication, assembly, and exit of single virus particles can be studied in real time in living cells. For example, the movement of HIV-1 particles labeled with GFP (by fusion with the viral protein Vpr) was beautifully visualized in cells that had been injected with rhodamine-tubulin to label microtubules.
Of course, virus particles are too small to have color. I once told this to a second-grade class as I was showing them the X-ray structures of eight different picornaviruses, each of which I had colored differently. One young man raised his hand and said “Do you make them different colors just to know which is which?” Who said you can’t teach virology in elementary school!
M Chalfie, Y Tu, G Euskirchen, W. Ward, D. Prasher (1994). Green fluorescent protein as a marker for gene expression Science, 263 (5148), 802-805 DOI: 10.1126/science.8303295
David C. Baulcombe, Sean Chapman, Simon Cruz (1995). Jellyfish green fluorescent protein as a reporter for virus infections The Plant Journal, 7 (6), 1045-1053 DOI: 10.1046/j.1365-313X.1995.07061045.x
K MORIYOSHI, L RICHARDS, C AKAZAWA, D OLEARY, S NAKANISHI (1996). Labeling Neural Cells Using Adenoviral Gene Transfer of Membrane-Targeted GFP Neuron, 16 (2), 255-260 DOI: 10.1016/S0896-6273(00)80044-6
Desai P Person S (1998) Incorporation of the green fluorescent protein into the herpes simplex virus type 1 capsid. J. Virol. 72:7563-7568.
M EKSTRAND (2008). The alpha-herpesviruses: molecular pathfinders in nervous system circuits Trends in Molecular Medicine, 14 (3), 134-140 DOI: 10.1016/j.molmed.2007.12.008
D. McDonald Vodicka MA, Lucero G, Svitkina TM, Borisy GG, Emerman M, Hope TJ. (2002). Visualization of the intracellular behavior of HIV in living cells The Journal of Cell Biology, 159 (3), 441-452 DOI: 10.1083/jcb.200203150

Very interesting post, I guess this blog become my primary information source about Virology , so much to learn and so much to read 🙂
Haha, cute. Start them young!
Eurokrize
Eesti-von Raivo Pommer–raimo1@hot.ee
L'euro poursuit son repli ce mardi face au dollar. Vers 18h45, un euro s'échangeait ainsi contre 1,32 dollar, après voir touché 1,3168 dollar, au plus bas depuis le 11 décembre. Lundi soir, un euro valait 1,3362 dollar. Les cambistes spéculent sur une probable baisse des taux européens à l'issue de la réunion du Conseil des gouverneurs de la Banque centrale européenne (BCE), ce jeudi à Francfort.
Face au ralentissement économique, l'institution présidée par Jean-Claude Trichet devrait opter pour un nouvel assouplissement monétaire. La majorité des économistes parient sur une baisse de 50 points de base du taux directeur européen, qui serait ainsi ramené à 2%.
La tendance baissière de la devise européenne est par ailleurs renforcée par les craintes sur la dette de plusieurs gouvernements de la zone euro après que l'agence de notation Standard & Poor's a placé la note de la dette à long terme de l'Etat espagnol sous surveillance négative. Cette dernière pourrait ainsi perdre son rang “AAA”.
De son côté, le billet vert a été soutenu par les propos de, Ben Bernanke. Le président de la Réserve fédérale américaine qui a estimé mardi que son institution disposait encore “d'outils puissants” contre la crise.
Pingback: Virus images at VIPERdb
too good information
very ineresting post about the picornaviruses. shout out to hop high.
there should be a video at youtube showing entry and exit of a virus !
There are videos of influenza and HIV-1 replication that are
reasonably good – see the links in notes for TWiV #60.
There are videos of influenza and HIV-1 replication that are
reasonably good – see the links in notes for TWiV #60.
Pingback: Even toy makers should get the science right
I am sorry but I have to ask. Why are viruses to small to have colour?
the GFP is green and is smaller than the virus. If colour is a measure of the energy of a photon then wouldn't a hydrogen atom be the smallest thing that could have colour in that it is the simplest system in which the change of its electron's energy state would emit a photon of a given energy (AKA colour)? I am mising something obvious here aren't I?
After thought
Are you just meaning it is so small that light in our visible range is too long in wave length to resolve it hence the need for electron rather than optical microscope? It would still have a colour we just can't 'see' it.
If a virus has a chromophore you would be able to see the color in a
concentrated solution (or with good microscope and a bright
fluorescent label) you can see the fluorescence from a single
particle. In this case you could actually locate it very precisely
(much more accurately than the wavelength of the light), but of course
could not resolve two viruses closer than the wavelength limit. In
the absence of a chromophore, you can generate color by scattering
light. Some viruses a very large, sufficiently large to scatter light
strongly. Crystals of tipula iridescent virus actually diffract light
creating rainbow patters. Solutions of these viruses would appear
irredescent (hence the name) and colors could vary depending on the
angle that you are looking at the solution, like an opal. Very
concentrated solutions of viruses as small as poliovirus, like the
band of virus in a CsCl gradient appear to be blue, due to
preferential scattering of the blue light, but this is almost
certainly due to the formation of large aggregates of varying sizes
approaching the wavelenghts off UV, and blue light. The electronic
transitions that you refer to in hydrogen would occur either at the
very long wavelength (radio) for rotations, or very short wavelengths
(x-ray) for true electronic transitions. Bond stretches occur in the
IR, visible light transitions always involve highly conjugated
systems. Hence the statement that if a virus had a visible light
chromophore solutions would appear to be colored. As indicated above
the other source of color is light scattering (which makes the sky
blue and the sunset red). The scattering of light requires particle
sizes that approach the wavelength of the light. Here most viruses
are too small to scatter visible light unless they are aggregating,
except the very large viruses whose diameters are either close to or
as large as the wavelength of visible light.
I am sorry but I have to ask. Why are viruses to small to have colour?
the GFP is green and is smaller than the virus. If colour is a measure of the energy of a photon then wouldn't a hydrogen atom be the smallest thing that could have colour in that it is the simplest system in which the change of its electron's energy state would emit a photon of a given energy (AKA colour)? I am mising something obvious here aren't I?
After thought
Are you just meaning it is so small that light in our visible range is too long in wave length to resolve it hence the need for electron rather than optical microscope? It would still have a colour we just can't 'see' it.
If a virus has a chromophore you would be able to see the color in a
concentrated solution (or with good microscope and a bright
fluorescent label) you can see the fluorescence from a single
particle. In this case you could actually locate it very precisely
(much more accurately than the wavelength of the light), but of course
could not resolve two viruses closer than the wavelength limit. In
the absence of a chromophore, you can generate color by scattering
light. Some viruses a very large, sufficiently large to scatter light
strongly. Crystals of tipula iridescent virus actually diffract light
creating rainbow patters. Solutions of these viruses would appear
irredescent (hence the name) and colors could vary depending on the
angle that you are looking at the solution, like an opal. Very
concentrated solutions of viruses as small as poliovirus, like the
band of virus in a CsCl gradient appear to be blue, due to
preferential scattering of the blue light, but this is almost
certainly due to the formation of large aggregates of varying sizes
approaching the wavelenghts off UV, and blue light. The electronic
transitions that you refer to in hydrogen would occur either at the
very long wavelength (radio) for rotations, or very short wavelengths
(x-ray) for true electronic transitions. Bond stretches occur in the
IR, visible light transitions always involve highly conjugated
systems. Hence the statement that if a virus had a visible light
chromophore solutions would appear to be colored. As indicated above
the other source of color is light scattering (which makes the sky
blue and the sunset red). The scattering of light requires particle
sizes that approach the wavelength of the light. Here most viruses
are too small to scatter visible light unless they are aggregating,
except the very large viruses whose diameters are either close to or
as large as the wavelength of visible light.
Viruses are the same color every single everything else is… absolute jet blackness. Other “colors” are optical illusions created by the human brain interpreting the different photon frequencies. Photons and the electrons that caused them to be admitted are both absolute jet blackness.
Typo alert: that should read “emitted” and not “admitted”.
How often is an electron microscope used to study and analyze a virus? Thanks.