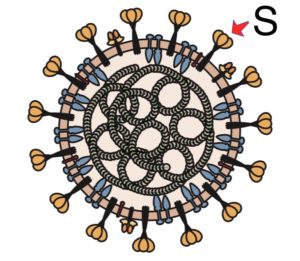

The membrane of coronaviruses harbors a trimeric transmembrane spike (S) glycoprotein (pictured) which is essential for entry of virus particles into the cell. The S protein contains two functional domains: a receptor binding domain, and a second domain which contains sequences that mediate fusion of the viral and cell membranes. The S glycoprotein must be cleaved by cell proteases to enable exposure of the fusion sequences and hence is needed for cell entry.
The nature of the cell protease that cleaves the S glycoprotein varies according to the coronavirus. The MERS-CoV S glycoprotein contains a furin cleavage site and is probably processed by these intracellular proteases during exit from the cell. The virus particles are therefore ready for entry into the next cell. In contrast, the SARS-CoV S glycoprotein is uncleaved upon virus release from cells; it is likely cleaved during virus entry into a cell.
Proteolytic cleavage of the S glycoprotein can determine whether the virus can cross species, e.g. from bats to humans. For example, the S glycoprotein from a MERS-like CoV from Ugandan bats can bind to human cells but cannot mediate virus entry. However, if the protease trypsin is included during infection, the S glycoprotein is cleaved and virus entry takes place. This observation demonstrates that cleavage of the S glycoprotein is a barrier to zoonotic coronavirus transmission.
Examination of the protein sequence of the S glycoprotein of SARS-CoV-2 reveals the presence of a furin cleavage sequence (PRRARS|V). The CoV with the highest nucleotide sequence homology, isolated from a bat in Yunnan in 2013 (RaTG-13), does not have the furin cleavage sequence. Because furin proteases are abundant in the respiratory tract, it is possible that SARS-CoV-2 S glycoprotein is cleaved upon exit from epithelial cells and consequently can efficiently infect other cells. In contrast, the highly related bat CoV RaTG-13 does not have the furin cleavage site.
Whether or not the furin cleavage site within the S glycoprotein of SARS-CoV-2 is actually cleaved remains to be determined. Meanwhile, it is possible that the insertion of a furin cleavage site allowed a bat CoV to gain the ability to infect humans. The furin cleavage site might have been acquired by recombination with another virus possessing that site. This event could have happened thousands of years ago, or weeks ago. Upon introduction into a human – likely in an outdoor meat market – the virus began its epidemic spread.
Furins are also known to control infection by avian influenza A viruses, in which cleavage of the HA glycoprotein is needed for entry into the cell. Low-pathogenic avian influenza viruses contain a single basic amino acid at the cleavage site in the HA protein which is cleaved by proteases that are restricted to the respiratory tract. Insertion of a furin cleavage site in the HA of highly pathogenic avian H5N1 influenza viruses leads to replication in multiple tissues and higher pathogenicity, due to the distribution of furins in multiple tissues.
Acquisition of the furin cleavage site might be viewed as a ‘gain of function€™ that enabled a bat CoV to jump into humans and begin its current epidemic spread.

This is fascinating, in particular the difference between SARS (2003) and SARS-CoV-2 (2019). I have a couple questions, but I’ll summarize what I understood first, in case I’m not following.
They explain that there are two necessary cleavages that must occur for infection. First, the S protein must be split into S1/S2, because those two sub-units mediate the distinct tasks of attachment and entry, respectively. Then the S2 must be cleaved at the “S2-prime” site, splitting the fusion peptide (FP) from the “internal fusion peptide” (IFP), because “it is likely both… participate in the viral entry process.” About the S2 cleaving, they say: “The furin-like S2′ cleavage site … is identical between the 2019-nCoV and SARS-CoV”.
About the S1/S2 cleaving they say that it “exhibits different motifs” among CoVs, and point to a “site1 & site2” on SARS-CoV-2. If I understand correctly, they say that site 2, which also appears on both SARS-CoV-2 and SARS-CoV, is known to be left uncleaved on viral egress and therefore must somehow be cleaved prior to viral attachment in the absence of another S1/S2 cleavage mechanism. “Site 1” is the new furin-like cleavage site, and it does not appear in SARS-CoV (they do say that other CoV, e.g., MERS-CoV) “harbor[] furin-like [S1/S2] cleavage sites”). This region on SARS-CoV-2 “contains 12 additional nucleotides upstream” to site 1, “which corresponds to a canonical furin-like cleavage site”.
If site 1 is cleaved on SARS-CoV-2 S1/S2 on viral exit, then there is no need for further cleaving on entry, so this is the potential “gain-of-function” for the new virus. This potential for an infectivity gain — due to the cleaving of S1/S2 on exit, as compared to the SARS necessity for cleaving on entry — seems to be the most important idea here. Obviously there are lots of factors that go into “infectivity” (and maybe I’m improperly associating within-host infectivity of cells with the observed epidemiological-scale infectivity), but if this effect is important, then have two questions:
(1) If MERS-CoV also had this “site 1” furin-like cleavage site, why did it have a similarly low infectivity to SARS?
(2) The “site 1” aa sequence in SARS-CoV-2019 is different than those of the other CoV possessing a furin-like cleavage “site 1”. Is it possible to find this specific sequence in another viruses, which could implicate them as a co-infection partner that led to this mutation?
The putative furin cleavage site as listed in the blog exists in both COVID-19 and bat strain RaTG13.
No this site is absent in the RaTG13 and in the pangolin sequence.
Our observations is also present in the excellent paper on the Spike cryo-EM from MCLeLLan lab.
“Cryo-EM Structure of the 2019-nCoV Spike in the Prefusion Conformation
Daniel Wrapp, Nianshuang Wang, Kizzmekia S Corbett, Jory A Goldsmith, Ching-Lin Hsieh, Olubukola Abiona, Barney S Graham, View ORCID ProfileJason S McLellan
doi: https://doi.org/10.1101/2020.02.11.944462
Ballistol is a powerfull anti viral product:
https://algerietouteheure.com/2020/02/12/coronavirus-ballistol /
One well known human coronavirus, HCoV-NL63 also uses ACE2 for cell entry.
Clinical spectrum looks like COVID19 :
“mild to moderate upper respiratory tract infections, severe lower respiratory tract infection.”
https://en.wikipedia.org/wiki/Human_coronavirus_NL63
So its not necessarily ACE2-affinity and the SARS-spike protein that makes the virus “new and dangerous” ?
Maybe COVID19 could (hopefully) be not that “SARS-like” sever after all ?
Maybe there is 10x more, but completely unreported very mild “common cold” type of COVID19 illness prevalence in China ?
Whats you virologists take on this ?
Sorry for the mistake I made in my alignment. PRRA is indeed unique to COVID-19. My question is how important the acquired cleavage site is for viral infection. If it is important for viral exit, then the released viral particles would have the truncated spike protein missing S1. If it is important for viral entry, how do you reconcile with the fact that other coronaviruses such as SARS would be able to infect without this site?
Another interesting and related paper was released today: “The Proximal Origin of SARS-CoV-2”, which also discusses the insertion of the furin cleavage site:
http://virological.org/t/the-proximal-origin-of-sars-cov-2/398
Someone named Bill Gallaher left a comment on that page about the similarity/differences between the RaTG13 bat coronavirus generally, and the furin-cleavage insertion specifically. I’d be interested to hear what others have to say about his comments and conclusions.
He points to an earlier post he made on the subject, 10 days ago:
http://virological.org/t/tackling-rumors-of-a-suspicious-origin-of-ncov2019/384
in which he compared the _nucleotide_ sequences of SARS-CoV-2 and RaTG13. Two things jump out in this comparison:
(1) Although the aa sequences of the two agree exactly over a segment of ~200 residues surrounding the insertion, the nucleotide agreement is only 93% (19 out of 288 are mutations). He mentions a rule of thumb that “wobble base mutagenesis occurs at a rate of 1% per decade”, suggesting a common ancestor circa 1950*.
(2) Although there is a 12 nucleotide insertion in the SARS-CoV-2, compared to RaTG13, it is offset from the aa code. With respect to the RaTG13 sequence, which translates to QTNSR|SV, the insertion is actually _within_ the Serine that precedes the cleavage point. The SARS-CoV-2 insertion in the nucleotide sequence actually looks like this:
UCA —-> U[CU CCU CGG CGG G]CA
where the first nucleotide of RaTG13’s Serine becomes the (alternatively spelled) Serine preceding the code for PRR in the SARS-CoV-2 insertion, and the last two nucleotides of RaTG13’s Serine complete the code for the final aa, Alanine, of the insertion**.
* — Or would it be 70/2 years ago, c1985, with each strain accounting for half of the disagreement?
** — I guess an alternative explanation would be that the UCA mutated to UCU, prior to an insertion (after RaTG13’s Serine) of [CCU CGG CGG GCA].
Furin was discovered in 1986. The first furin inhibitor was patented in 1994. Here is a patent review published in 2015 including 17 patents for inhibitors, primarily, but also for novel uses of furin itself.
https://www.tandfonline.com/doi/full/10.1517/13543776.2014.1000303?src=recsys&
The authors include a cautionary remark about the specific problem discussed here:
“…using furin [inhibition] against viruses which utilize glycoproteins with a furin cleavage site may not always produce beneficial effects, as revealed by the Ebola virus gB protein …[even though] a furin cleavage site has been clearly demonstrated, blocking the furin-mediated cleavage of the Ebola virus gB does not result in a reduction of viral replication…†Suggests Ebola may have another trimmer.
Furin inhibitors have been studied for their antiviral effects but clinical trial data is limited.
So far I have not been able to identify any FDA approved furin inhibitors. Furin is a vital enzyme with hundreds of known substrates, so perhaps this is not surprising.
Bloomberg has reported that in addition to approved drugs, candidates for clinical trials in China include TCM’s (traditional Chinese medicines). Google turned up a TCM whose active ingredient is identified in the following link as a furin inhibitor: baicalein.
http://www.eurekaselect.com/106526/article
I will look for confirmation from other sources that baicalein is a furin inhibitor. There are papers about baicalein as an antiviral, notably against dengue. Baicalein (5,6,7-trihydroxyflavone) is a flavone, originally isolated from the roots of Scutellaria baicalensis Georgi (Huang Qin in Chinese), which has a long history of practice in TCM. It is also reported in Oroxylum indicum or Indian trumpet flower and Scutellaria lateriflora.
John
Furin was discovered in 1986. The first furin inhibitor was patented in 1994. Here is a link to a patent review published in 2015 including 17 patents, most of them for furin inhibitors.
https://www.tandfonline.com/doi/full/10.1517/13543776.2014.1000303?src=recsys&
The authors include a cautionary remark about the specific problem discussed here:
“…using furin [inhibition] against viruses which utilize glycoproteins with a furin cleavage site may not always produce beneficial effects, as revealed by the Ebola virus gB protein …[even though] a furin cleavage site has been clearly demonstrated, blocking the furin-mediated cleavage of the Ebola virus gB does not result in a reduction of viral replication…â€
Suggests Ebola has another cutter.
Historically furin inhibitors have been studied for their antiviral effects but clinical trial data is very limited, and so far I have not found any FDA approved furin inhibitors. Furin is a vital enzyme with hundreds of known substrates, so perhaps this is not surprising.
Bloomberg has reported that in addition to approved drugs, candidates for clinical trials in China include TCM’s (traditional Chinese medicines). Google turned up a TCM whose active ingredient is identified in the following link as a furin inhibitor: baicalein.
http://www.eurekaselect.com/106526/article
I will look for confirmation from other sources that baicalein is a furin inhibitor. There are papers about baicalein as an antiviral, notably against dengue. Baicalein (5,6,7-trihydroxyflavone) is a flavone, originally isolated from the roots of Scutellaria baicalensis Georgi (Huang Qin in Chinese), which has a long history of practice in TCM. It is also extracted from Oroxylum indicum or Indian trumpet flower and Scutellaria lateriflora.
“Sequence Analysis Indicates that 2019-nCoV Virus Contains a Putative Furin Cleavage Site at the Boundary of S1 and S2 Domains of Spike Protein”
https://osf.io/nkcrf
“a furin cleavage sequence (PRRARS|V)” should be PRRAR|SV. i believe that this is just a typo.
“Upon introduction into a human – likely in an outdoor meat market – the virus began its epidemic spread.”
Please stop spreading unfounded rumors. (This goes both ways!) There is no evidence that the meat market was the origin of the outbreak. Neither were the first cases related to the market (Huang, Lancet, 2020) nor were any animals found that could have been the source.
Pingback: Pangolins and the origin of SARS-CoV-2 coronavirus
E. Braun, D. Sauter. Furin-mediated protein processing in infectious diseases and cancer, August, 2019.
https://onlinelibrary.wiley.com/doi/pdf/10.1002/cti2.1073
Current review of therapeutic inhibition of furin.
A search turned up four furin inhibitors that have been identified and extracted from traditional medicinal plants in China and India. Some have been tested for antiviral activity against Dengue. All four are flavonoids.
The appeal of plants and extracts with a history of use in traditional medicines is that there is experience – sometimes long experience — in observing their impacts on human patients. Few of the perhaps 150 agents that have been identified as furin inhibitors have ever been in clinical trials.
In 2010 at the University of Ottawa, three flavonoid furin inhibitors were extracted from the traditional medicinal plant, Oroxylum Indicum: baicalein, chrysin and oroxylin. Oroxylin was the most potent inhibitor of furin with a k sub i of 5 µM. All three inhibit other PCs in addition to furin.
https://www.researchgate.net/profile/Ajoy_Basak2/publication/43347486_Proprotein_Convertase_Inhibitory_Activities_of_Flavonoids_Isolated_from_Oroxylum_Indicum/links/55d14dc208ae6a881385ec40/Proprotein-Convertase-Inhibitory-Activities-of-Flavonoids-Isolated-from-Oroxylum-Indicum.pdf
Luteolin is another flavonoid with furin inhibitory activity. It is found in many types of plants including medicinal herbs. It was tested as a furin inhibitor for its antiviral activity against Dengue. The K sub i was in the high fifties, as recounted here:
https://www.ncbi.nlm.nih.gov/pubmed/28389141
Reply to Carston. The Huang et al (2020) paper in the Lancet makes EXPLICIT reference to the seafood market.
“We report the epidemiological, clinical, laboratory, and radiological characteristics, treatment, and clinical outcomes of 41 laboratory-confirmed cases infected with 2019-nCoV. 27 (66%) of 41 patients had a history of direct exposure to the Huanan seafood market.”
Pingback: TWiV 588: Coronavirus update - Save the pangolin! | This Week in Virology
I’m doing an animation that will show how we currently think cell entry works for COVID-19. Is it safe to say we currently think endocytosis is needed (as was reported for SARS), or might this furin site allow direct fusion at the cell membrane?
Here’s the paper claiming endocytosis was needed for SARS: https://www.nature.com/articles/cr200815
Hello everyone. I have no idea what I’m reading here but it all sounds very well thought through. I am just a paranoid germaphobe and a mother of 5 young children trying to figure out if I should use be stressed about the coronavirus thing. I don’t trust the news and everything I find online directs me to nbc, fox, or the cdc. I don’t even know how I came across this article. Any prevention tips or things I should look for? The front page of our local paper says they are investigating a case in my county. I am slightly freaking out.
Pingback: SARS-CoV-2 LÂY LAN MẠNH NHẤT, VÀ COVID-19 SẼ LÀ ÄẠI DỊCH ! | Tran Ba Thoai's Blog
@Calla
“Some coronavirus advice I’ve gleaned from folks who’ve worked in or studied other epidemics. Please take seriously and pass on to family and friends…”
https://twitter.com/alimanfoo/status/1232693018295250946
Pingback: COVID-19, Furins & Hypoxia - The Vitamin C Connection - EvolutaMente.it
On cleavage mechanism of SARS-CoV-2 spike see recent paper https://www.researchsquare.com/article/086968bf-2235-499d-a33c-3615ff20bc7f/v1
“Second, we found that 2019-nCoV S protein mediated entry on 293/hACE2 cells
was mainly through endocytosis, and PIKfyve, TPC2, and cathepsin L are critical for virus entry. ”
So Cathepsin L inhibitors may be effective in treatment SARS-CoV-2.
Interestingly, the bovine milk proteins possesses such property:
https://www.ncbi.nlm.nih.gov/pubmed/12788072
“New functions of lactoferrin and beta-casein in mammalian milk as cysteine protease inhibitors.
We found new inhibitory function of lactoferrin and beta-casein in milk against cysteine proteases using reverse zymography. The inhibition of cathepsin L by lactoferrin was strongest and the inhibition kinetics were of a non-competitive type.”
Unfortunately, adult chinese peoples do not drink bovine milk due to lactose intolerance.
Pingback: SARS-CoV-2 lây lan mạnh nhất và COVID-19 sẽ là đại dịch – Kim Dung/Kỳ Duyên
@Jon Perry
Did you ever find out any additional information regarding whether endocytosis is needed? I’m trying to determine if the interest in Baricitinib is warranted due to AAK1 inhibition disrupting endocytosis. My search lead me to your comment on this blog, but I haven’t found any answers yet.
Thanks,
Mark
Pingback: ÐÐ¿Ð¸Ð´ÐµÐ¼Ð¸Ñ ÐºÐ¾Ñ€Ð¾Ð½Ð°Ð²Ð¸Ñ€ÑƒÑа перераÑтает в пандемию • «Похудела Я» - Сайт пр
I would like to ask for your opinion on a hypothesis I have formulated regarding the SARS-Cov-2 viral entry routes. I understand the 2003 SARS virus infects cells by using the Spike Protein (“S”) to bind with the ACE2 receptor protein. However the ACE2 receptor is not present in many cells in healthy individuals, which could explain the “relatively low” contagion rate of the 2003 SARS virus.
Your article and the following article posted on South China Morning Post which
references a pre-print paper, published on Chinaxiv.org, by a team from Nankai University, shows that a cleavage in the spike protein allows it to use the furin protein to allow a “direct fusion” of the viral and cellular membranes.
https://www.scmp.com/news/china/society/article/3052495/coronavirus-far-more-likely-sars-bond-human-cells-scientists-say
Since furin is a ubiquitous protein, the infection rates would be 100 to 1000 greater than the 2003 SARS CoV.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1964754/
My hypothesis is that SARS-Cov-2 follows a two phase infection route:
1. In the first phase it infects cells through the furin protein, thanks to
its spike protein cleavage. This would account for its relatively high infection rate
and initially mild symptoms. It could also explain why asymptomatic patients shed the virus.
2. In a second phase, once in the body, the SARS-Cov-2 virus would infect cells through the traditional 2003 SARS route, namely the ACE2 receptors in lower lung
cells, which would result in the severe pneumonia and 2003 SARS-like symptoms.
I would be very happy if you could share this hypothesis with your virology colleagues and tell me what your think. It may have important implications
for vaccine development and therapeutics research.
Thank you kindly
Pingback: Protecting ourselves from Covid-19? – The Garden Clinic
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2583654/
Entry from the Cell Surface of Severe Acute Respiratory Syndrome Coronavirus with Cleaved S Protein as Revealed by Pseudotype Virus Bearing Cleaved S Proteinâ–¿
Rie Watanabe,1 Shutoku Matsuyama,1 Kazuya Shirato,1 Masami Maejima,1 Shuetsu Fukushi,2 Shigeru Morikawa,2 and Fumihiro Taguchi1,*
Departments of Virology III,1 I, National Institute of Infectious Diseases, Murayama Branch, Musashi-Murayama, Tokyo 208-0011, Japan2
Dec. 2008
Severe acute respiratory syndrome (SARS) coronavirus (SARS-CoV) is known to take an endosomal pathway for cell entry; however, it is thought to enter directly from the cell surface when a receptor-bound virion spike (S) protein is affected by trypsin, which induces cleavage of the S protein and activates its fusion potential. This suggests that SARS-CoV bearing a cleaved form of the S protein can enter cells directly from the cell surface without trypsin treatment. To explore this possibility, WE INTRODUCED A FURIN-LIKE CLEAVAGE SEQUENCE in the S protein at amino acids 798 to 801 and found that the mutated S protein was cleaved and induced cell fusion without trypsin treatment when expressed on the cell surface. Furthermore, a pseudotype virus bearing a cleaved S protein was revealed to infect cells in the presence of a lysosomotropic agent as well as a protease inhibitor, both of which are known to block SARS-CoV infection via an endosome, whereas the infection of pseudotypes with an uncleaved, wild-type S protein was blocked by these agents. A heptad repeat peptide, derived from a SARS-CoV S protein that is known to efficiently block infections from the cell surface, blocked the infection by a pseudotype with a cleaved S protein but not that with an uncleaved S protein. Those results indicate that SARS-CoV with a cleaved S protein is able to enter cells directly from the cell surface and agree with the previous observation of the protease-mediated cell surface entry of SARS-CoV. …
https://www.researchgate.net/publication/276359777_A_Single_Point_Mutation_Creating_a_Furin_Cleavage_Site_in_the_Spike_Protein_Renders_Porcine_Epidemic_Diarrhea_Coronavirus_Trypsin_Independent_for_Cell_Entry_and_Fusion (2015)
The emerging porcine epidemic diarrhea [corona] virus (PEDV) requires trypsin supplementation to activate its S protein for membrane fusion and virus propagation in cell culture. By substitution of a single amino acid in the S protein WE CREATED A RECOMBINANT PEDV (PEDV-S(FCS)) WITH AN ARTIFICIAL FURIN PROTEASE CLEAVAGE SITE N-terminal of the putative fusion peptide. PEDV-S(FCS) exhibited trypsin-independent cell-cell fusion and was able to replicate in culture cells independent of trypsin, though to low titer. …
Lots of “gain of function†research going on everywhere. And synthetic biology is all the rage!
——
https://www.edge.org/conversation/william_mcewan-molecular-cut-and-paste
William McEwan: This afternoon I received in the post a slim FedEx envelope containing four small vials of DNA. The DNA had been synthesized according to my instructions in under three weeks, at a cost of 39 U.S. cents per base pair (the rungs adenine-thymine or guanine-cytosine in the DNA ladder). The 10 micrograms I ordered are dried, flaky, and barely visible to the naked eye, yet once I have restored them in water and made an RNA copy of this template, THEY WILL ENCODE A VIRUS I HAVE DESIGNED. …
PRRARS is always mentioned as a furin cleavage site.
PRRARV turns out to be a textbook fibronectin adhesion molecule (heparin binding sequence). This in turn seems to be used by some viruses for infection. Any ideas on this?
“The DNA had been synthesized according to my instructions in under three weeks, at a cost of 39 U.S. cents per base pair (the rungs adenine-thymine or guanine-cytosine in the DNA ladder). The 10 micrograms I ordered are dried, flaky, and barely visible to the naked eye, yet once I have restored them in water and made an RNA copy of this template, THEY WILL ENCODE A VIRUS I HAVE DESIGNED. …”
Yep, it seems like ISIS won’t have to bother trying to get nukes or drones anymore… bio warfare is all the rage now. Even Iran seems to be already going at it full stop, looking at the numbers of Iranian individuals spreading 2019-Ncov across Middle east… a cheap way to put the West to their knees.
@Pete
Interesting question. Indeed, there are some evidence that the PRRARV sequence motif in fibronectin may be utilized for macrophage and/or virus adhesion. Here PRRARV is at the host cell side. On the other hand, the PRRARSV sequence motif at the S1 subunit and S2 subunit boundary of spike protein of SARS0-CoV-2 virus is the sequence motif for host cell surface protease – Furin to recognize and to cleave. There is no conflict.
PRRARS is just one of the possible Furin cleavage sequence motifs. Furin recognize what some called ‘polybasic cleavage site’. The most pronounced feature is P1 and P4 being Arg residue (ie. RXYR, where X or Y should be Arg or Lys).
Â
Â
Pingback: COVID-19 jako SARS-CoV-2 z mutacjÄ… glikoproteiny S zapobieganie i zwalczanie | Marzena Kolano - Porady dietetyczne, Dieta, Suplementy - Infinite Progress Nutrition
Potential furin inhibitors:
Skullcap (Oroxylin A, Chrysin, Luteolin)
Thyme, Sage, Artichoke (Luteolin)
Andrographis paniculata (Andrographolid)
https://www.ncbi.nlm.nih.gov/pubmed/23231348https://sci-hub.se/https://www.ncbi.nlm.nih.gov/pubmed/23835774https://www.ncbi.nlm.nih.gov/pubmed/22716122 https://www.ncbi.nlm.nih.gov/pubmed/22642118
Pingback: KHÃC BIỆT GIá»®A SARS-CoV-2 VỚI VIRUS CÚM | Tran Ba Thoai's Blog
I posted the following comments on another page of the site without any answer so far. I am very happy to see here other comments like mine. I am a microbiologist with experience creating mutants (not viruses) and I know that we have all the methods to manipulate genomic sequences without leaving any trace..
My previous comments:
What about producing a chimeric virus combing KP876546 or RaTG13 (or similar) with part of the RBD from CoVs isolated from pangolins (or similar) searching the missing link between bats and humans? Furin cleavage site might come in addition to enhance virulence.
In this work a possible procedure to generate chimeric viruses is described:
Manipulation of the Coronavirus Genome Using Targeted RNA Recombination with Interspecies Chimeric Coronaviruses
Cornelis A.M. de Haan, Bert Jan Haijema, Paul S. Masters, and Peter J.M. Rottier
Another interesting information:
Title: Methods and compositions for chimeric coronavirus spike proteins United States Patent 9884895 Inventors: Baric, Ralph (Haw River, NC, US) Agnihothram, Sudhakar (Ellicott City, MD, US) Yount, Boyd (Hillsborough, NC, US) Publication Date: 02/06/2018
Assignee:
The University of North Carolina at Chapel Hill (Chapel Hill, NC, US)
I have now a question: why the famous RaTG13 sequence, from a sample collected in 2013 was submitted to NCBI only on 27-JAN-2020, after the outbreak of SARS-CoV-2?
From NCBI:
LOCUS MN996532 29855 bp RNA linear VRL 24-FEB-2020
DEFINITION Bat coronavirus RaTG13, complete genome.
ACCESSION MN996532
VERSION MN996532.1
KEYWORDS .
SOURCE Bat coronavirus RaTG13
ORGANISM Bat coronavirus RaTG13
Viruses; Riboviria; Nidovirales; Cornidovirineae; Coronaviridae;
unclassified Coronaviridae.
REFERENCE 1 (bases 1 to 29855)
AUTHORS Zhu,Y., Yu,P., Li,B., Hu,B., Si,H.R., Yang,X.L., Zhou,P. and
Shi,Z.L.
TITLE Direct Submission
JOURNAL Submitted (27-JAN-2020) CAS Key Laboratory of Special Pathogens,
Wuhan Institute of Virology, Center for Biosafety Mega-Science,
Chinese Academy of Sciences, No. 44 Xiao Hong Shan, Wuhan, Hubei
430071, China
source 1..29855
/organism=â€Bat coronavirus RaTG13″
/mol_type=â€genomic RNAâ€
/isolate=â€RaTG13″
/isolation_source=â€fecal swabâ€
/host=â€Rhinolophus affinisâ€
/db_xref=â€taxon:2709072″
/country=â€Chinaâ€
/collection_date=â€24-Jul-2013″
Are some virologist scared that the true will come out and their research will be limited? I am ready to take the risk to see also my work limited. Playing with the fire is dangerous without precautions…
Pingback: “Corona Virus is Bioweapon” – www.spash.co.ke
Pingback: Social Media Says Scientists May Prevent COVID-19 Spread-Updates
Info on Furin/PCs and virus cell entry mechanism
http://dx.doi.org/10.3390/v11090837
https://www.mdpi.com/1999-4915/11/9/837
@Rossana
Not sure if you have read “The Proximal Origin of SARS-CoV-2” by Kristain G. Andersen et al. http://virological.org/t/the-proximal-origin-of-sars-cov-2/398
If not certainly it’s worth reading. This article expressed the view that it’s very unlikely that SARS-COV-2 was engineered. Any thoughts?
@gwo Sure that I read that article. It gave me the idea that SARS-CoV 2 could have been genetically manipulated starting from a virus similar to RaTG13 (is this sequence to be trusted completely? Is it published in some articles?) and combining it with a CoV sequence from another host, for example pangolin. Check Fig. 1: If you switch the aa sequence in the RBD from the pangolin with the sequence from RaTG13 the resulting sequence is almost identical to SARS-CoV2.
…Both the polybasic cleavage site and O-linked glycans are unique to SARS-CoV-2 and not previously seen in lineage B betacoronaviruses.
This can come also from a lab, maybe even more easily than from natural recombination. There are published works on the manipulation of the polybasic cleavage site in coronaviruses.
..Further, if genetic manipulation had been performed, one would expect that one of the several reverse genetic systems available for betacoronaviruses would have been used. However, this is not the case as the genetic data shows that SARS-CoV-2 is not derived from any previously used virus backbone.
Sure that all the sequences obtained from bat or other animal screening have been deposited? Why RaTG13, isolated in 2013, has been made public only in 2020?
…However, the acquisition of the polybasic cleavage site or O-linked glycans – if functional – argues against this scenario. New polybasic cleavage sites have only been observed after prolonged passaging of low pathogenicity avian influenza virus in cell culture or animals.
This can be easily done in a lab as well. See PMID: 24667706 and PMID: 30209269.
..The generation of SARS-CoV-2 by cell culture or animal passage would have required prior isolation of a progenitor virus with a very high genetic similarity.
Same as before. Can you exclude that such kind of virus has been isolated and not made public because it was under study?
https://www.nature.com/articles/d41586-020-00660-x
Nature reporter’s sampling of research and opinions on the furin cleavage site as of March 6, with a correction on March 11. Also includes a list of references associated with investigators who offered comments.
Pingback: COVID-19, Pneumonia & Inflammasomes - The Melatonin Connection - EvolutaMente.it
Pingback: Furin cleavage site in the SARS-CoV-2 coronavirus glycoprotein – Virology Hub