
The new study, a collaboration among scientists at the Food and Drug Administration, the National Institutes of Health, and Harvard Medical School, utilized samples from 37 CFS patients obtained in the mid-1990s. A key difference from earlier studies is that some repeat samples were used: four obtained two years later and frozen, and eight taken in 2010 and processed without freezing. The patients were from New England, New York State, and North Carolina and were not part of the previous study in which XMRV was detected. Control samples were obtained from 44 healthy blood donors.
Two kinds of polymerase chain reaction (PCR) assays were used to determine the presence of XMRV. In the first, DNA was extracted from PBMC to search for the presence of proviral DNA (e.g., viral DNA integrated into the host chromosome). In the second, RNA was extracted from plasma, and subjected to reverse-transcriptase PCR to detect viral RNA.
The authors found that 32 of 37 samples from CFS patients (86.5%) were positive in the PCR assay of PBMC DNA, compared with 3 of 44 samples (6.8%) from healthy controls. All four samples taken 2 years later, and 7 of the 8 samples taken 15 years later from CFS patients were also positive in this assay. The latter is an important result because it shows presence of virus in patients for long periods. Viral RNA was detected in plasma of 42% of the samples.
The authors performed additional experiments to rule out false positive results. The nucleotide sequences of all PCR products were determined to ensure that they represented the expected target sequence. Furthermore, contamination with mouse DNA was excluded by assaying for murine mitochondrial DNA – it was not found in any of the samples tested.
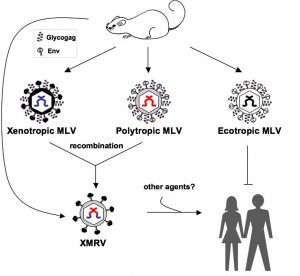
Perhaps the most interesting finding, aside from the detection of viral RNA in a large fraction of CFS patients, is the observation that the virus detected is not XMRV. Nucleotide sequence analysis of PCR amplified DNA revealed 96.6% nucleotide identity with XMRV, but they are clearly different viruses. The RNA genome of XMRV is a recombinant between the genomes of xenotropic* and polytropic* murine leukemia viruses (Figure). The genome of the viruses detected in the new study appears to be derived from polytropic MLVs, and could be called PMRV. This observation indicates that both xenotropic and polytropic MLV can infect humans in North America.
What is the significace of the observation that both XMRV and PMRV have been associated with CFS? Some investigators have suggested that variation of XMRV is minimal, based on the close identity of sequences obtained from prostate cancer and CFS patients. The finding of a different virus, PMRV, suggests greater genetic variation than previously thought, and could in part explain previous failures to detect MLV-related viruses in the previous four studies of CFS patients. Other explanations, including geographic differences and patient heterogeneity, are also plausible.
I find it very interesting that the authors readily detected PMRV sequences in both PBMC DNA and plasma. The authors of the 2009 Science paper have suggested that these approaches are not sufficiently sensitive to detect viral nucleic acid, and that culturing with susceptible cells must be done to amplify the virus. That explanation, which was used to explain the failure to detect XMRV in the previous four studies, is not likely to be correct. The authors of those studies might want to repeat their assays using primers more likely to detect PMRV.
Where do these findings lead next? The authors of an accompanying commentary have two appropriate suggestions:
At this juncture, it would seem reasonable to conduct extensive case-control studies in North America…using coded samples from subject with inflammatory disease to determine the frequency of MLV infection in patients with CFS. The potential transmission of MLV-related sequences from human to human should also be epidemiologically evaluated. […]it would also be appropriate to conduct interventional studies.
It is possible that these studies, which are needed to determine if MLVs cause human disease, might not have been done without a second independent confirmation of the association of MLV-related viruses with CFS.
*Xenotropic murine leukemia viruses infect non-mouse cells, while polytropic murine leukemia viruses can infect the cells of mice and other mammals.
Lo SC, Pripuzova N, Li B, Komaroff AL, Hung GC, Wang R, & Alter HJ (2010). Detection of MLV-related virus gene sequences in blood of patients with chronic fatigue syndrome and healthy blood donors. Proceedings of the National Academy of Sciences of the United States of America, 107 (36), 15874-9 PMID: 20798047

What does PBMC stand for? I don’t look as this as a right or wrong, but an opportunity to venture into the viral world further than ever before. I suspect there will be a definitive answer soon and obviously a very important one for all of us that will be logging into “Not CFSCentral” and “Phoenix Has Risen” someday to share stories of having a life!!! My gut feeling is that the results of these studies are going to cause a period of evolution in the understanding of virology and the treatment and cure of numerous chronic diseases. This will go far beyond Prostate Cancer and CFS!!
Thanks for continuing to follow this story.
1. It’s interesting that when Alter/Lo sequenced their PCR products, they found that there were 3 PMRV variants that CFS patients had but that the 3 positive products from the controls did not fit into these three categories. In terms of geographic variation I wonder if the CFS subjects from a particular area were more likely to have a common CFS type (e.g. those from NY more likely to have type 2) and also if the variation in the controls might be related to the fact that they were drawn from DC blood banks rather than blood banks from NY, New England, or NC (where subjects were drawn from).
2. In your above quote from the commentary, this was confusing to me:
“At this juncture, it would seem reasonable to conduct extensive case-control studies in North America, as suggested by Lo et al. (6), using coded control samples from subjects with inflammatory disease to determine the frequency of MLV infection in patients with CFS. ”
Why would they pick control subjects with inflammatory disorders in particular?
3. Has anyone thought about examining the mice in areas where CFS outbreaks have occurred?
Pingback: Tweets that mention PMRV joins XMRV as possible etiologic agent of chronic fatigue syndrome -- Topsy.com
Given the relative short sequences (http://www.ncbi.nlm.nih.gov/nuccore?term=mlv-related%20virus%20cfs) made by Lo/Atler, do we know enough to say these findings are, or not, XMRV?
Professor, thank you for this concise and elegant analysis.
Did you attend the workshop in Maryland this week? Can you give us your analysis of those proceedings as well?
Professor,
Thanks for the post and the chance to comment. There are three suggestions in the quoted passage (not two), so it’s not quite clear whether you are endorsing interventional studies being begun now. While opposition to such studies is not incomprehensible to me, I strongly support them. In particular, the most interesting such study on CFS, to date, has been Dr John Chia’s work on the use of interferon-alpha and ribavirin. This paper, which may or may not be the only one on the subject, was published in 2004:
http://www.jarcet.com/articles/Vol4Iss2/Chia2-Jar-spring.pdf
In fact, not only is this worthy of attention now, but it is moreover puzzling that there has not already been a follow-on study at some point during the last six years, unless of course there is some grave epistemologic objection to Chia’s efforts.
You can see the spectacular nature of the improvement, generally followed by relapse starting a few weeks after withdrawal of IFNa. This is, of course, quite interesting with reference to a potential retroviral etiology, since Type I Interferons are robustly active in retroviral infections, though certainly also in other viral infections. It is also interesting to consider that the characteristic relapse might (or might not) be preventable by the right sort of lifelong small-molecule anti-retroviral treatment.
We definitely know that the sequenced parts are not XMRV. It's conceivable that the other parts (yet unsequenced) could have come from XMRV in a recombination (essentially, 'hybridization') event. But even then, the whole thing could not really be termed 'XMRV'.
Pingback: My Chronic Fatigue Syndrome Healing and Remission Secrets
Very interesting, thank you. Could it be that we infect people with PMRVs through the blood supply?
Fair enough, but I’ll defer to Dr. Coffin from the Q&A session on the matter if these sequences constitute new viruses. We need complete sequences for that. Dr. Coffin said “And so I think clearly a bunch of different sequences. I think before we start referring to a bunch of different viruses we really need to have viruses that go with those sequences. We can say there are sequences there that look like endogenous MLV’s, maybe a little bit different from them. But we just don’t have a virus yet to go with them. “
Many thanks for your continued coverage of the latest developments with these viruses, Dr. Racaniello.
These new findings are really interesting.
The presence of very strong positive MLV (PMRV)-specific bands in plasma also caught my eye.
You wonder about these readily detected PMRV sequences in both PBMC DNA and plasma, because authors of the 2009 Science paper had suggested that these approaches are not sufficiently sensitive to detect viral nucleic acid, and that culturing with susceptible cells must be done to amplify the virus.
The authors of the Science paper, however, have suggested this after the paper was published. Nowhere in the Science paper is any mentioning whatsoever of culturing as a step before PCR. Have you seen their extremely strong signals on the gel after one single round of PCR?
They only published recently in an addendum to the Science paper in a new virology journal (?!), that PCR is the least sensitive method.
Another reason for the bands being strong in plasma (stronger then in PBMC), may be that there are less “competing” nucleic acids than in plasma than in PBMC, were the low frequency virus nucleic acids are amidst the abundant nucleic acids from blood cells.
What also struck me was that Lo et al also used different inner primers vs Lombardi et al. These inner primers were more MLV-conservative. I looked at them more carefully and noticed 2 mismatches with XMRV in one of these inner primers. I really wonder whether this would at least partially explain why others (including Lombardi et al) didn't pick up any PMRV. It doesn't explain however, that Lo et al didn't find any XMRV. ( Lo et al also used the XMRV-specific inner primers, only in their article they are not clear when they use what)
Other remarkable findings: (1) much more heterogeneity in MLV-sequences than reported by Lombardi and (2) the control MLV's (only 3 or so) seem to differ from the patient MLV's.
I find it highly unlikely that the samples examined by Lo et al find contain 100% PMRV, and those of Lombardi et al 100% XMRV. As a matter of fact the WPI (of the Science paper) now also mentions the presence of MLV in their samples. I don't know whether they have changed the primers they used.
From Twitter I understood that one conclusion from the 1st workshop on XMRV was that conditions could make the difference. I tend to believe the same (and populations/classification of CFS might explain differences if good quality PCR-assays using PBMC-enriched fresh blood samples fail to demonstrate anything)
I'm very curious as to your opinion about this.
This is my last blog post on XMRV discussing this:
http://laikaspoetnik.wordpress.com/2010/08/30/does-the-nhifda-paper-confirm-xmrv-in-cfs-well-ditch-the-mr-and-scratch-the-x-and-youve-got-mlv/
Jacqueline
While the focus of this blog is peer reviewed research it was interesting to note that Judy Mikovits made a curious remark at the recent XMRV workshop.
“we have learned literally in past 2 wks in the BWG (blood working group which includes the WPI, NIH, CDC etc.) that processing may be a key and we may have found an opportunity to have a processing protocol where everyone would at least find viral RNA in plasma and blood products without culture.”
Dr. Mikovits has a tendency to make off the cuff remarks, but it will be interesting to see if this grows legs.
Are you saying that this PMRV is NOT in the MLV Family that was recently suggested as growing that XMRV was also indeed a part of?
This seems confusing esp since the following was included in this post:
“Where do these findings lead next? The authors of an accompanying commentary have two appropriate suggestions:
At this juncture, it would seem reasonable to conduct extensive case-control studies in North America…using coded samples from subject with inflammatory disease to determine the frequency of MLV infection in patients with CFS. The potential transmission of MLV-related sequences from human to human should also be epidemiologically evaluated. […]it would also be appropriate to conduct interventional studies.”
That's a nice diagram at the beginning of this article, could PLease explain WHERE on that would PMRV fit?
If PMRV also fits in the MLV family that can lead to XMRV,
would PMRV just be like another sibling in the same MLV category?
So if you test +Positive for MLV… then you would STILL know
that you could have either XMRV or PMRV ?
Please clarify……… and Thanks..
wheres Dr Goff these days!! He's a fab researcher and would love to see him do some talks again.
Dr Goff is one of our favourites in the xmrv story.
I did not attend the 1st XMRV workshop – but there is a terrific summary at http://www.cfids.org/xmrv/wrkshp-report.pdf
I spoke with Dr. Coffin at ICAAC (I wanted to interview him for TWiV but he could not make it). He said that the Lo/Alter isolates 'are not XMRV; they are not the same virus'. He is correct that we don't yet have infectious virus, but he seems convinced that they are different.
Yes, it's possible. That's why the blood supply is being examined very carefully.
PBMC: Peripheral blood mononuclear cell such as a lymphocyte, monocyte or macrophage. Very easy to purify from total blood.
Dr. Goff is at a meeting in Venice this week. We'll definitely bring him back to TWiV – after all, he's in the office next to mine and easy to speak with.
Pingback: Is XMRV a laboratory contaminant?
Pingback: Authors retract paper on detection of murine leukemia virus-releated sequences in CFS patients