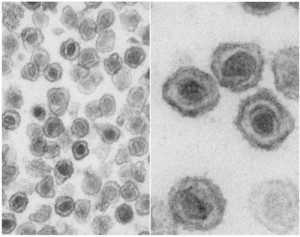

Polymerase chain reaction was used to detect XMRV in 267 respiratory samples taken from German patients. One group comprised sputum and nasal swab specimens from 75 travelers from Asia who had respiratory tract infections. The second group consisted of 31 bronchoalveolar lavage samples from patients with chronic obstructive pulmonary disease, while samples from the third group were from 161 immunosuppressed patients with severe respiratory tract infections. The study included 62 healthy controls. It should be noted that none of the patients had been diagnosed with CFS.
XMRV sequences were detected in 3 of 75 samples (2.3%) in group 1, 1 of 31 samples (3.2%) in group 2, and 16/161 (9.9%) in group 3. Six of the XMRV-positive samples in the second group also contained rhinovirus, adenovirus, or pathogenic fungi. The higher rate of detection of XMRV and other microbes in immunosuppressed individuals is not unexpected. The control group contained 2 of 62 samples (3.2%) positive for XMRV.
The presence of XMRV in PBMCs and plasma suggests a blood-borne route of transmission of the virus: transfusions, health care associated needle sticks, and intravenous drug use. Does finding XMRV in the respiratory tract prove that the virus can be transmitted by the respiratory route? No, not until we have other information, including the level of virus in respiratory secretions, and the infectivity of XMRV. In this context it is interesting to note that it was not possible to isolate infectious XMRV from the respiratory tract of the German patients.
Reviewing the transmission of another human retrovirus, HIV-1, is instructive in understanding the pathogenesis of XMRV infection. The main modes of transmission of HIV-1 are sexual, parenteral, and from mother to infant. These routes of transmission are consistent with levels of infectious virus in body fluids (shown in this table). Viral RNA can be detected at several levels of the respiratory tract, but respiratory secretions rarely transmit HIV.
FIsher, N., Schulz, C., Stieler, K., Hohn, O., Lange, C., Drosten, C., & Aepfelbacher, M. (2010). Xenotropic murine leukemia virus-related gammaretrovirus in respiratory tract Emerg. Inf. Dis. : 10.3201/eid1606.100066

Hi there.
Is there anything here which might indicate why previous German studies have been unable to find any XMRV?
We seem to be getting a lot of variable results with CFS and prostate cancer, and no-one seems to know why.
If this new paper is accurate, would that indicate that the CFS papers which found 0% of patients had XMRV were probably unable to detect XMRV infection?
Germany detected XMRV in over 2% of the public, this fits in line with thousands of miles away in USA (WPI SCIENCE study of October 2009), and also in a Japanese study.
Failure to detect XMRV is down to method. PCR is unreliable, and the method used to detect XMRV has to work to be able to detect it. We already know that the UK Wessely & McClure study was incapable of detecting XMRV (via the method used) and the Dutch XMRV CFS study again also.
Only 1 'failed' XMRV CFS study exists where the Scientists have no history of anti CFS public rhetoric, and that is the XMRV Kerr study, again in the UK. The two aforementioned chose CFS people without 'symptoms of organic disease' and also had a cohort of 'tired' peoplea attending psychiatric clinics. ME and CFS are not classified as psychiatric diseases, hence the obvious concerns of this 'choice' of cohort. Having said that, the cohort is not of paramount importance but the methods used to detect XMRV.
Politics in XMRV and CFS is rife. Wessely & McClure UK XMRV study, the patients were hand picked by Professor Simon Wessely from the UK, who has a vested interest to halt XMRV research in UK CFS patients as soon as possible. (Simon Wessely is on record calling ME CFS patients 'neurotic', 'disgusting', 'free from blame and guilt'). Other than offensive rhetoric, it is important to note Wessely has also stated ''Viral attribution (reflects) somatization par excellance''. It is therefore utterly unsurprising to see that Wessely & McClure 'failed' to find XMRV in CFS.
The fact XMRV has been detected in the healthy populace at rates of around 2 – 4% in 3 different countries separated by thousands of miles, says a lot. It's out there and lab contamination is not an issue. Method is everything, as all scientists know and they also know politics should stay out of science also as it pollutes research. Lastly, there is more than one strain of XMRV, and the prostate cancer strain is not the same as found in CFS.
“would that indicate that the CFS papers which found 0% of patients had XMRV were probably unable to detect XMRV infection?”
I'm no scientist, but I'm following rigorously these studies and my answer to you is that I think it would absolutley prove that the studies that didn't find XMRV in ME/CFS patients – at least if their cohort had a significant number of ME/CFS patients, and not just patients who suffer from fatigue because of depression or something else – were not able to find XMRV even if it's there.
The prove is that after at least 2 studies being done in Germany about XMRV in prostate cancer, that didn't find XMRV in prostate tissue, here is a study in the same country that finds it in respiratory secretions – and that is because even though they used just PCR, they used it correctly this time.
Here's the breakdown on XMRV/prostate cancer studies so far; they are really all over the map-
Negative XMRV/prostate cancer studies-
-German prostate cancer study-
0/589 German prostate cancer patients XMRV+
'Lack of evidence for xenotropic murine leukemia virus-related virus(XMRV) in German prostate cancer patients'
Hohn O, Krause H, Barbarotto P, Niederstadt L, Beimforde N, Denner J, Miller K, Kurth R, Bannert N.Retrovirology. 2009 Oct 16;6:92.
http://www.ncbi.nlm.nih.gov/pubmed/19835577
-Irish prostate cancer study-
0/139 Irish prostate cancer patients XMRV+
'NO EVIDENCE OF XMRV IN IRISH PROSTATE CANCER PATIENTS WITH THE R462Q MUTATION'
D'Arcy F., Foley R., Perry A., Marignol L., Lawler M., Gaffney E., Watson R.G.W., Fitzpatrick J.M., Lynch T.H.
European Urology Supplements Volume 7, Issue 3, Page 271 (March 2008)
http://www.europeanurology-supplement.com/artic…
– 'Failure to detect XMRV in human prostate tumors' – poster presentation at upcoming Cold Springs Harbor Retroviruses Conference 2010
Aloia, A.L.
http://meetings.cshl.edu/meetings/abstracts/201…
———————————-
Positive XMRV/prostate cancer studies-
-XMRV/prostate cancer study #1(US)-
8/20 prostate cancer patients XMRV 1/66 (1.5%) different kind of prostate cancer patients XMRV
'Identification of a novel Gammaretrovirus in prostate tumors of patients homozygous for R462Q RNASEL variant'
Urisman A, Molinaro RJ, Fischer N, Plummer SJ, Casey G, Klein EA, Malathi K, Magi-Galluzzi C, Tubbs RR, Ganem D, Silverman RH, DeRisi JL.
PLoS Pathog. 2006 Mar;2(3):e25. Epub 2006 Mar 31.
http://www.ncbi.nlm.nih.gov/pubmed/16609730
-XMRV/prostate cancer study #2(US)-
334 consecutive prostate resection specimens, XMRV DNA found in 6% and XMRV protein expression in 23% of prostate cancers vs. 6/101 of controls(6%) XMRV+
'XMRV is present in malignant prostatic epithelium and is associated with prostate cancer, especially high-grade tumors'
Schlaberg R, Choe DJ, Brown KR, Thaker HM, Singh IR.
Proc Natl Acad Sci U S A. 2009 Sep 22;106(38):16351-6.
http://www.ncbi.nlm.nih.gov/pubmed/19805305
XMRV/prostate cancer study #3(US)-
At a serum dilution of 1:150, our assay detected 11 (27.5%) of 40 patients with XMRV neutralizing antibodies, including 8 (40%) of 20 with the RNASEL genotype QQ and 3 (15%) of 20 with either the RQ or RR genotype. These results were in complete concordance with 2 other assays (polymerase chain reaction and fluorescence in situ hybridization), which were designed to detect XMRV infection
'XMRV infection in patients with prostate cancer: novel serologic assay and correlation with PCR and FISH'
Arnold RS, Makarova NV, Osunkoya AO, Suppiah S, Scott TA, Johnson NA, Bhosle SM, Liotta D, Hunter E, Marshall FF, Ly H, Molinaro RJ, Blackwell JL, Petros JA.
Urology. 2010 Apr;75(4):755-61.
http://www.ncbi.nlm.nih.gov/pubmed/20371060
In an interview on Australian radio, UK virologist Myra McClure stated that 'we're actually finding it[XMRV]' in prostate cancer samples, although how much is unclear at present.
http://www.abc.net.au/rn/healthreport/stories/2…
——————————
Equivocal XMRV/prostate cancer studies-
-German prostate cancer study #2-
1/105 non-familial prostate cancer patients and 1/70 tissue samples from men without prostate cancer XMRV+
'Prevalence of human gammaretrovirus XMRV in sporadic prostate cancer'
Fischer N, Hellwinkel O, Schulz C, Chun FK, Huland H, Aepfelbacher M, Schlomm T.J
Clin Virol. 2008 Nov;43(3):277-83.
http://www.ncbi.nlm.nih.gov/pubmed/18823818
-CDC prostate cancer study- (presented at CROI 2010)
Of 165 prostate tissues, 2 (1.2%) were positive by pol and env polymerase chain reaction
'Prevalence of Xenotropic Murine Leukemia Virus in Prostate Cancer'
William Switzer*, H Jia, H Q Zheng, S Tang, and W Heneine
CDC, Atlanta, GA, US
http://www.retroconference.org/2010/Abstracts/3…
***Note- Plasma from both persons (5956 and 6203) were negative by reverse transcriptase-polymerase chain reaction indicating absence of viremia. Both patients were Western blot-negative…The finding of undetectable antibodies and viremia in 2 patients is noteworthy and may reflect sequestered or cleared infections
———————————–
So three positive studies, two negative studies, two equivocal studies, one more unpublished negative study and one possibly positive unpublished study just in prostate cancer studies, which are still going on 4 years after the initial report of the virus in prostate cancer samples. Yeesh.
I have learn much from this article >:D< .Thank you so much
“Lastly, there is more than one strain of XMRV, and the prostate cancer strain is not the same as found in CFS”
Link please?
From what I have read, XMRV is a simple gammaretrovirus, that is very stable (also a reason why it might be able to survive in saliva, and this theory would speak to why infants and children under 15 years of age contracted the illness, form someone else that was in the acute infectious phase of it. Yes, this is all just theory, it will all be sorted out in due time). I have not read anything concerning strain differences, but would love a link to any intel that you reported above.
It's worthy to note that the failed CFS/XMRV studies were not replication studies. And unless Any XMRV association with CFS study is done PRECISELY the way it was done byt the WPI, CC and the NCI, then they most likely will fail. I supose they could call what they did verificaiton studies, but the cohorts and the assays used did no way, shape or form follow what was done in the original study.
NOTE: The Netherland study failed to note that there were positive for XMRV samples found by WPI in the samples used by the Netherlands. They kinda left that part out.. yeah.
There are now several phylogenetic trees that relate strains to geography… i don't think there is any clear link to specific strains causing one disease over another… i don't think there is a prostate cancer xmrv strain vs. xmrv… i think you most likely have different human phenotypes interacting with a highly similar cluster of retroviruses that has morphed as it has moved around the globe…. right now these trees are being used to track or show insight into the recent evolution of the virus not specify a disease etiology
If you go to http://www.retroconference.org/2010/data/files/… and click on 'friday' and scroll about 2/5 of the way down, that's where the XMRV presentations start. The CDC's presentation, 'Prevalence of Xenotropic Murine Leukemia Virus in Prostate Cancer' by Walid Heneine CDC, Atlanta, GA, US, shows some slides of phylogenic trees which show a different strain. The slides in question are 192, 193 and 194.
Jud, I may be able to add to your comment about whether there is a clear link to specific strains causing one disease over another. There are 6 strains of XMRV that have been isolated (and published):
FROM THE ORIGINAL SCIENCE ARTICLE: SUPPLEMENTAL MATERIALS:
“CFS XMRV strains WPI-1130, WPI-1138 and WPI-1169 in comparison to XMRV strains VP35, VP42 and VP62 derived from prostate cancer patients.”
From: http://www.anapsid.org/cnd/xmrv/xmrv-science-23…
Some more interesting info on the strains…
FROM BARANIUK'S ARTICLE ON XMRV IN CFS AND PROSTATE CANCER
“The six strains of XMRV that have been sequenced have greater than 99% identity, indicating
a new human infection rather than laboratory contamination.”
From: http://www.springerlink.com/content/416lq773u35…
FROM ANOTHER DISCUSSION FORUM
“If you compare the isolates that they had from the 3 prostate cancer cases, where they had actually cloned these, you can see, if you compare it to the reference strain, known as VP62, that’s the reference strain of what this virus looks like, the CFS samples here were clearly different, but they were highly similar – 99.7% – there were maybe 8 bases different across the entire 8,000 base pairs.”
From: http://forums.aboutmecfs.org/content.php?34-Tra…
and
“I read the excellent article with Proff Goff. Bearing in mind that there are at least six strains of XMRV one of his sentences hit me like a club between the eyes. He said that you only have to get your pcr primer wrong by only two nucleotides and you would not find the virus you were looking for if it was a different strain.”
FROM “CFS: COULD A STEALTH VIRUS BE LURKING
Here is what the (Science) paper reports. In 101 banked samples of PBMCs, 67% (68) were positive for a XMRV gag sequence. Next, seven of 11 PBMC CFS samples held at the Cleveland Clinic were shown to have XMRV gag plus env. Only 3.7% of PBMC DNA from healthy controls had XMRV gag when tested by PCR. Amazingly, those gag and env sequences were nearly identical to those from XMRV from prostate cancer-associated strains (PLoS Pathol. 2006;2:211). Full-length SMRV from two patients differed from prostate cancer strain VP62 by only six nucleotides, showing again a > 99% identity between the CFS and prostate cancer XMRV.
From: http://ahcpub.com/hot_topics/?htid=1&httid=2005
NOVEMBER 2009 HOT TOPICS
And finally (and I apologize for not being able to find this reference)… the researchers have noted that there is less variation between the CFS and prostate strains, than across the prostate strains themselves.
Another interesting twist is that while ME/CFS has been maligned in the research, interestingly several of the eminent prostate cancer researchers are crossing over to the dark side. Including Dr. Silverman, co-discovererer of XMRV in prostate cancer. And Dr Singh, who is not only researching XMRV in prostate cancer and ME/CFS, but also has forged ahead as you know, investigating the efficacy of antiretrovirals against XMRV in vitro. Smart, inquisitive researchers, open to the possibility that prostate cancer and Chronic Fatigue Syndrome may be unusual bedfellows with XMRV.
i suspect many will be looking into this if for no other reason than increased funding…
Pingback: XMRV prompts media thought: ask for the “state of play†| Code for Life